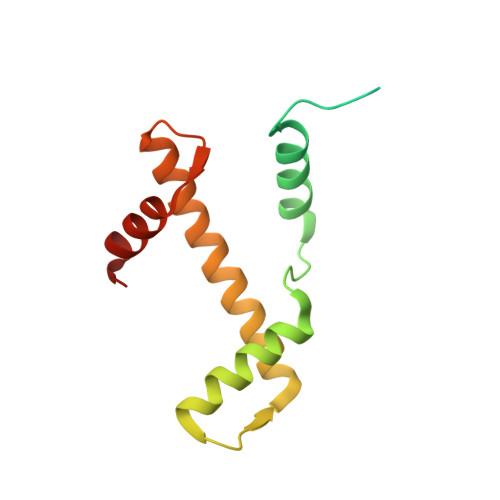
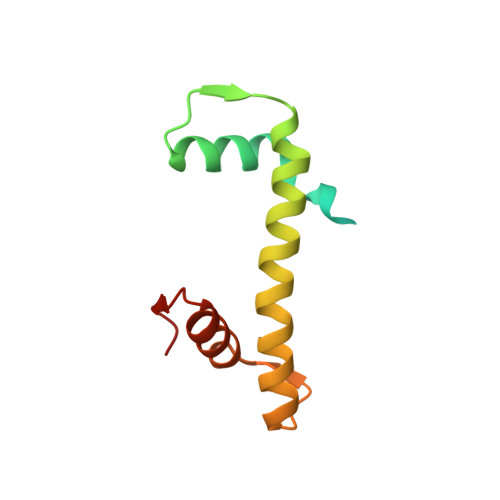
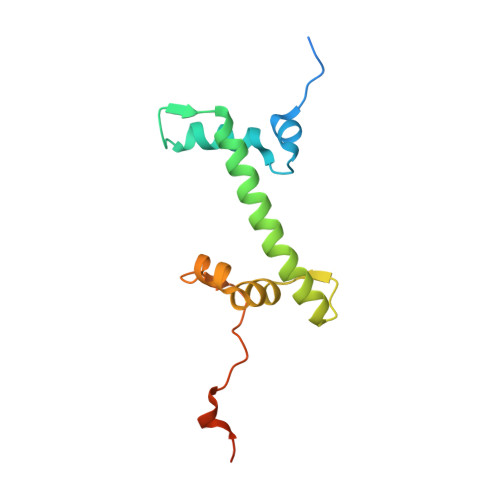
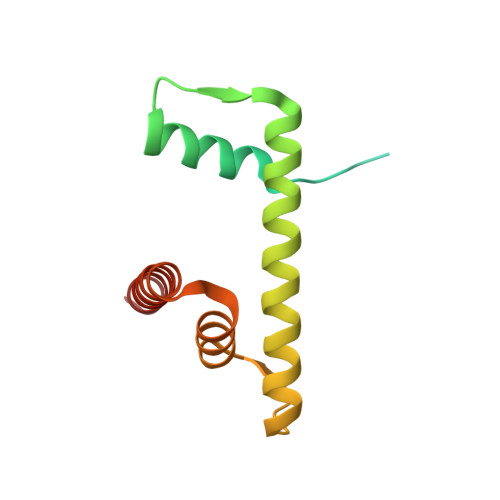
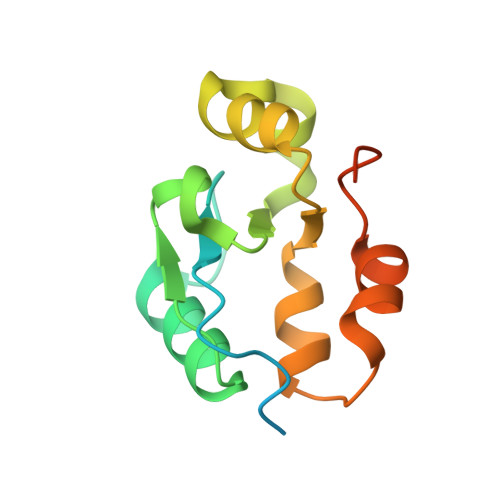
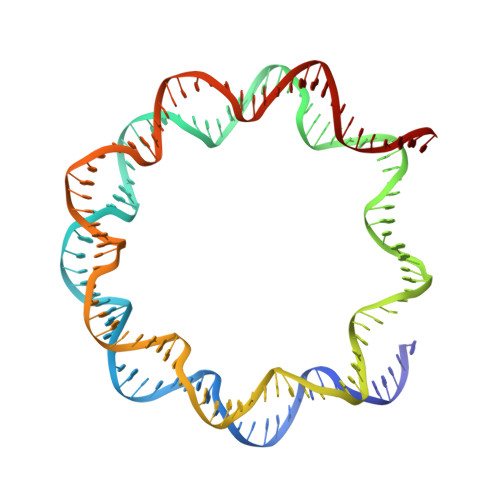
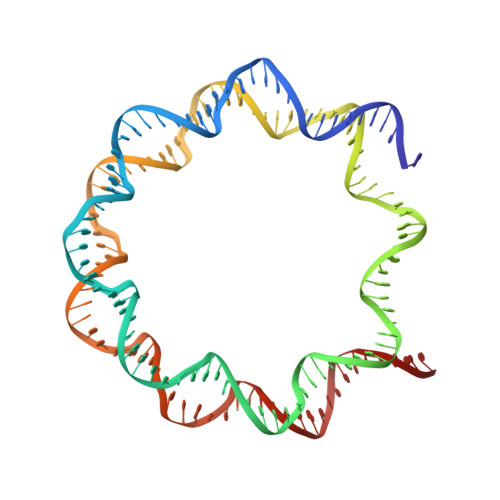
The BRCT domain of PARP1 binds intact DNA and mediates intrastrand transfer.
Rudolph, J., Muthurajan, U.M., Palacio, M., Mahadevan, J., Roberts, G., Erbse, A.H., Dyer, P.N., Luger, K.(2021) Mol Cell 81: 4994-5006.e5
- PubMed: 34919819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2021.11.014
- Primary Citation Related Structures:
7SCY, 7SCZ - PubMed Abstract:
PARP1 is a key player in the response to DNA damage and is the target of clinical inhibitors for the treatment of cancers. Binding of PARP1 to damaged DNA leads to activation wherein PARP1 uses NAD + to add chains of poly(ADP-ribose) onto itself and other nuclear proteins. PARP1 also binds abundantly to intact DNA and chromatin, where it remains enzymatically inactive. We show that intact DNA makes contacts with the PARP1 BRCT domain, which was not previously recognized as a DNA-binding domain. This binding mode does not result in the concomitant reorganization and activation of the catalytic domain. We visualize the BRCT domain bound to nucleosomal DNA by cryogenic electron microscopy and identify a key motif conserved from ancestral BRCT domains for binding phosphates on DNA and phospho-peptides. Finally, we demonstrate that the DNA-binding properties of the BRCT domain contribute to the "monkey-bar mechanism" that mediates DNA transfer of PARP1.
- Department of Biochemistry, University of Colorado Boulder, Boulder, CO 80309, USA.
Organizational Affiliation: