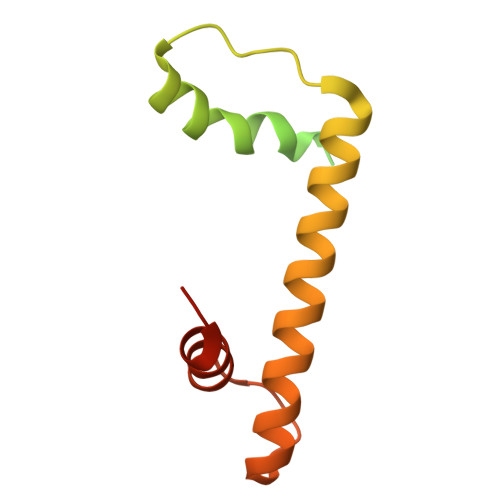
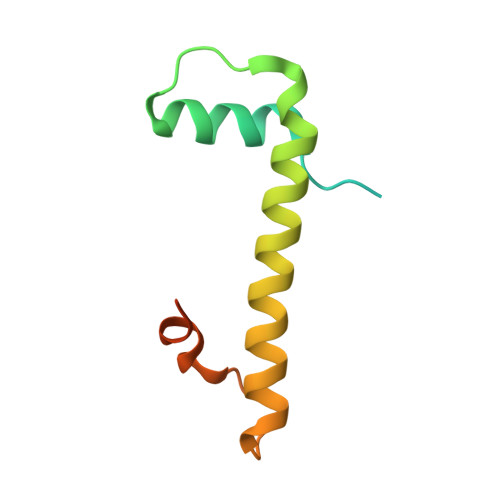
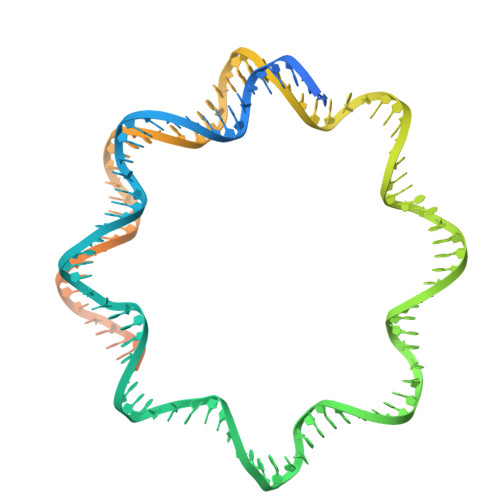
Non-nucleosomal (CENP-A/H4) 2 - DNA complexes as a possible platform for centromere organization.
Ali-Ahmad, A., Mors, M., Carrer, M., Li, X., Bilokapic, S., Halic, M., Cascella, M., Sekulic, N.(2025) bioRxiv
- PubMed: 39803555 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2024.12.31.630874
- Primary Citation Related Structures:
9GXA - PubMed Abstract:
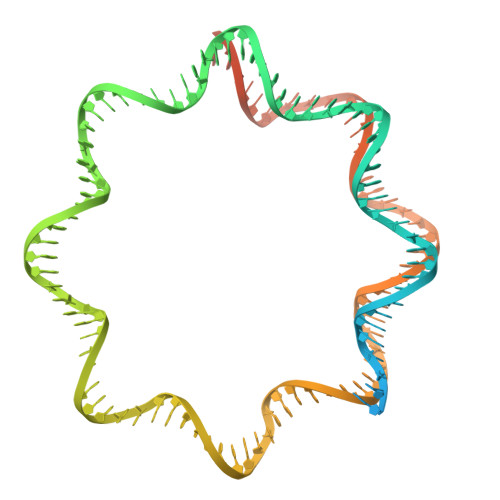
The centromere is a part of the chromosome that is essential for the even segregation of duplicated chromosomes during cell division. It is epigenetically defined by the presence of the histone H3 variant CENP-A. CENP-A associates specifically with a group of 16 proteins that form the centromere-associated network of proteins (CCAN). In mitosis, the kinetochore forms on the CCAN to connect the duplicated chromosomes to the microtubules protruding from the cell poles. Previous studies have shown that CENP-A replaces H3 in nucleosomes, and recently the structures of CENP-A-containing nucleosomes in complex with CCANs have been revealed, but they show only a limited interaction between CCANs and CENP-A. Here, we report the cryoEM structure of 2x(CENP-A/H4) 2 -di-tetramers assembled on DNA in the absence of H2A/H2B histone dimer and speculate how (CENP-A/H4) 2 -tetramers and -di-tetramers might serve as a platform for CCAN organization.
- Centre for Molecular Medicine Norway (NCMM), Nordic EMBL Partnership, Faculty of Medicine, University of Oslo, Oslo 0318, Norway.
Organizational Affiliation: