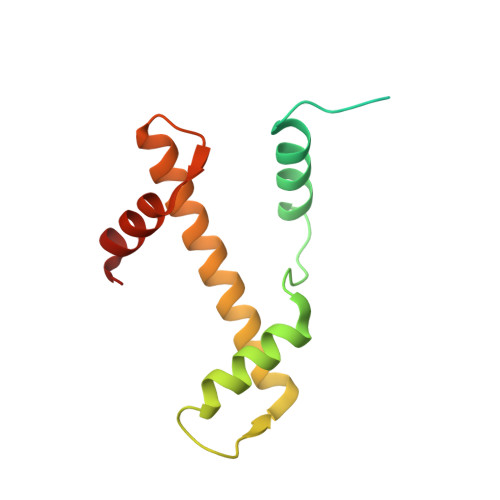
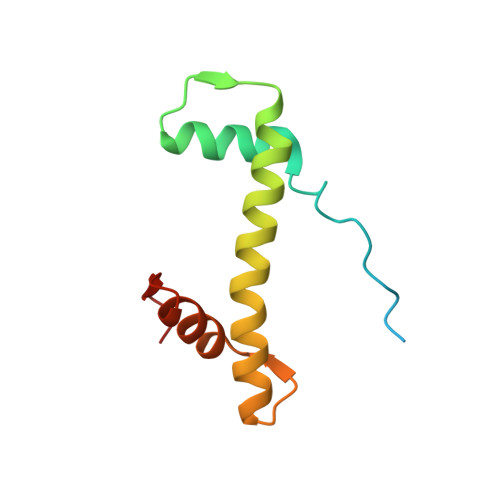
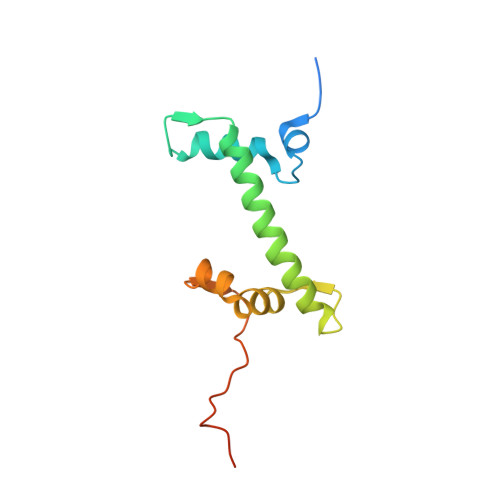
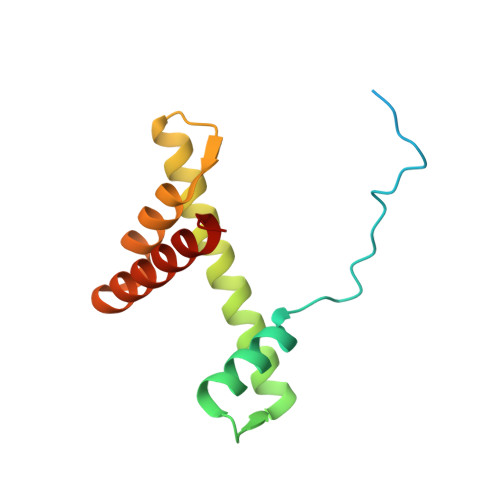
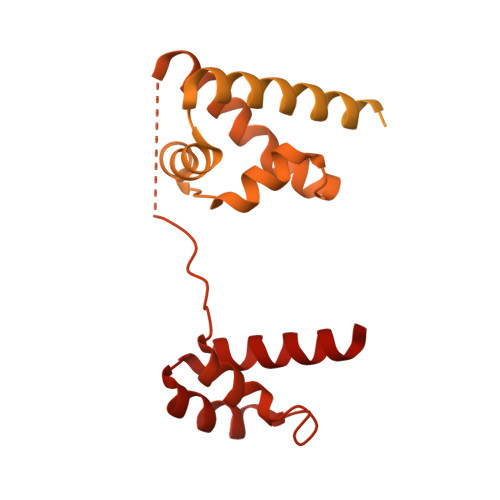
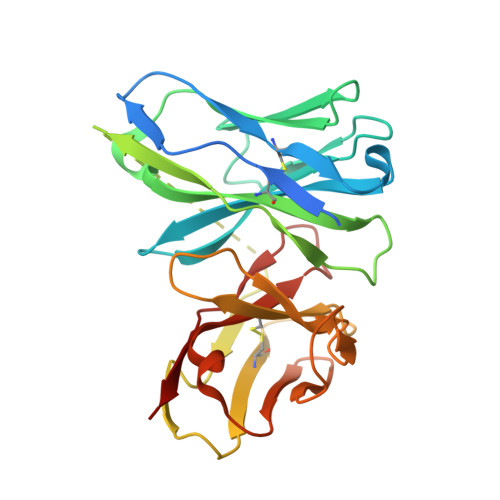
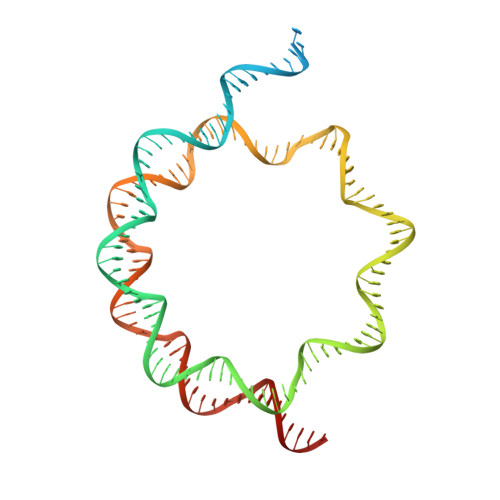
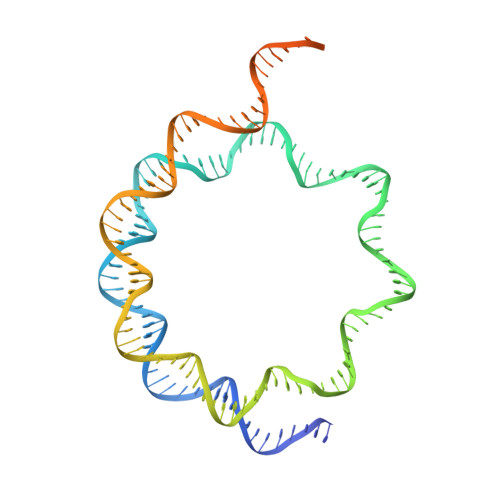
Structural mechanism of LIN28B nucleosome targeting by OCT4.
Guan, R., Lian, T., Zhou, B.R., Wheeler, D., Bai, Y.(2023) Mol Cell 83: 1970-1982.e6
- PubMed: 37327775 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2023.05.030
- Primary Citation Related Structures:
7U0G, 7U0I, 7U0J, 8DK5, 8SPS, 8SPU - PubMed Abstract:
Pioneer transcription factors are essential for cell fate changes by targeting closed chromatin. OCT4 is a crucial pioneer factor that can induce cell reprogramming. However, the structural basis of how pioneer factors recognize the in vivo nucleosomal DNA targets is unknown. Here, we determine the high-resolution structures of the nucleosome containing human LIN28B DNA and its complexes with the OCT4 DNA binding region. Three OCT4s bind the pre-positioned nucleosome by recognizing non-canonical DNA sequences. Two use their POUS domains while the other uses the POUS-loop-POUHD region; POUHD serves as a wedge to unwrap ∼25 base pair DNA. Our analysis of previous genomic data and determination of the ESRRB-nucleosome-OCT4 structure confirmed the generality of these structural features. Moreover, biochemical studies suggest that multiple OCT4s cooperatively open the H1-condensed nucleosome array containing the LIN28B nucleosome. Thus, our study suggests a mechanism of how OCT4 can target the nucleosome and open closed chromatin.
- Laboratory of Biochemistry and Molecular Biology, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: