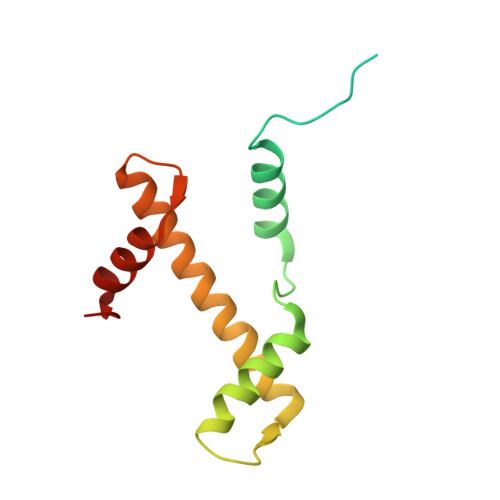
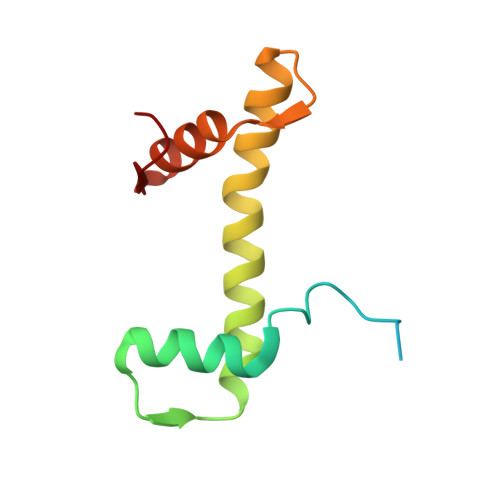
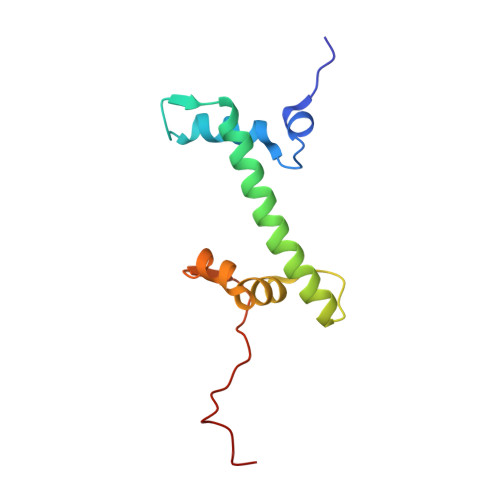
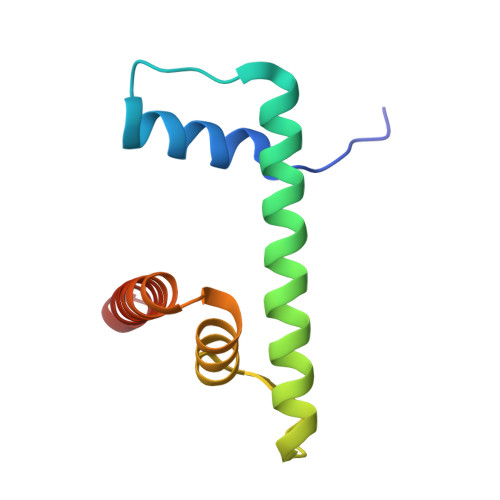
Structural basis for the inhibition of PRC2 by active transcription histone posttranslational modifications.
Cookis, T., Lydecker, A., Sauer, P., Kasinath, V., Nogales, E.(2025) Nat Struct Mol Biol 32: 393-404
- PubMed: 39774834 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01452-x
- Primary Citation Related Structures:
8VMI, 8VMJ, 8VML, 8VMN, 8VNV, 8VNZ, 8VO0, 8VOB - PubMed Abstract:
Polycomb repressive complex 2 (PRC2) trimethylates histone H3 on K27 (H3K27me3) leading to gene silencing that is essential for embryonic development and maintenance of cell identity. PRC2 is regulated by protein cofactors and their crosstalk with histone modifications. Trimethylated histone H3 on K4 (H3K4me3) and K36 (H3K36me3) localize to sites of active transcription and inhibit PRC2 activity through unknown mechanisms. Using cryo-electron microscopy, we reveal that histone H3 tails containing H3K36me3 engage poorly with PRC2 and preclude its effective interaction with chromatin, while H3K4me3 binds to the allosteric site in the EED subunit, acting as an antagonist that competes with activators required for spreading of the H3K27me3 repressive mark. Thus, the location of the H3K4me3 and H3K36me3 modifications along the H3 tail allows them to target two requirements for efficient trimethylation of H3K27 by PRC2. We further show that the JARID2 cofactor modulates PRC2 activity in the presence of these histone modifications.
- Department of Molecular and Cell Biology, University of California, Berkeley, CA, USA.
Organizational Affiliation: