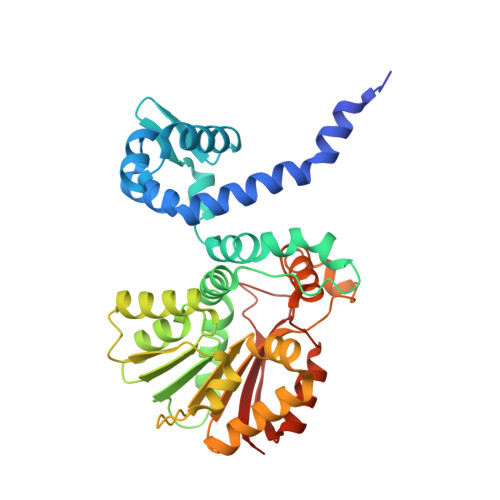
Structures and Protein Engineering of the alpha-Keto Acid C-Methyltransferases SgvM and MrsA for Rational Substrate Transfer.
Sommer-Kamann, C., Breiltgens, J., Zou, Z., Gerhardt, S., Saleem-Batcha, R., Kemper, F., Einsle, O., Andexer, J.N., Muller, M.(2024) Chembiochem 25: e202400258-e202400258
- PubMed: 38887142 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.202400258
- Primary Citation Related Structures:
8R4Z, 8RPR, 8RVC, 8RVS, 8RWM, 8RWW, 8RXF, 8RXG - PubMed Abstract:
S‑adenosyl-l-methionine-dependent methyltransferases (MTs) are involved in the C-methylation of a variety of natural products. The MTs SgvM from Streptomyces griseoviridis and MrsA from Pseudomonas syringae pv. syringae catalyze the methylation of the β-carbon atom of α-keto acids in the biosynthesis of the antibiotic natural products viridogrisein and 3‑methylarginine, respectively. MrsA shows high substrate selectivity for 5‑guanidino-2-oxovalerate, while other α-keto acids, such as the SgvM substrates 4-methyl-2-oxovalerate, 2-oxovalerate, and phenylpyruvate, are not accepted. Here we report the crystal structures of SgvM and MrsA in the apo form bound with substrate or S‑adenosyl-l-methionine. By investigating key residues for substrate recognition in the active sites of both enzymes and engineering MrsA by site-directed mutagenesis, the substrate range of MrsA was extended to accept α‑keto acid substrates of SgvM with uncharged and lipophilic β‑residues. Our results showcase the transfer of the substrate scope of α-keto acid MTs from different biosynthetic pathways by rational design.
- University of Freiburg, Institute of pharmaceutical science, GERMANY.
Organizational Affiliation: