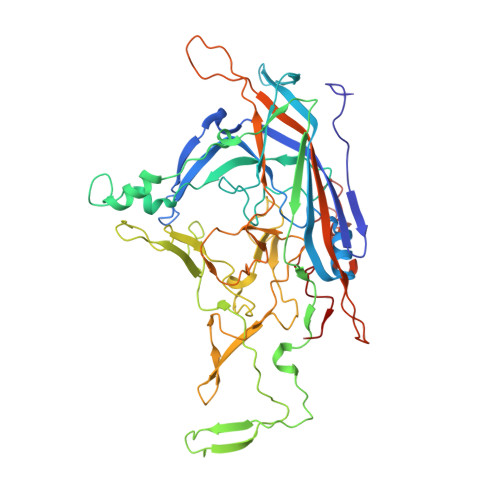
Characterization of the GBoV1 Capsid and Its Antibody Interactions.
Yu, J.C., Mietzsch, M., Singh, A., Jimenez Ybargollin, A., Kailasan, S., Chipman, P., Bhattacharya, N., Fakhiri, J., Grimm, D., Kapoor, A., Kucinskaite-Kodze, I., Zvirbliene, A., Soderlund-Venermo, M., McKenna, R., Agbandje-McKenna, M.(2021) Viruses 13
- PubMed: 33672786 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/v13020330
- Primary Citation Related Structures:
7LNK - PubMed Abstract:
Human bocavirus 1 (HBoV1) has gained attention as a gene delivery vector with its ability to infect polarized human airway epithelia and 5.5 kb genome packaging capacity. Gorilla bocavirus 1 (GBoV1) VP3 shares 86% amino acid sequence identity with HBoV1 but has better transduction efficiency in several human cell types. Here, we report the capsid structure of GBoV1 determined to 2.76 Å resolution using cryo-electron microscopy (cryo-EM) and its interaction with mouse monoclonal antibodies (mAbs) and human sera. GBoV1 shares capsid surface morphologies with other parvoviruses, with a channel at the 5-fold symmetry axis, protrusions surrounding the 3-fold axis and a depression at the 2-fold axis. A 2/5-fold wall separates the 2-fold and 5-fold axes. Compared to HBoV1, differences are localized to the 3-fold protrusions. Consistently, native dot immunoblots and cryo-EM showed cross-reactivity and binding, respectively, by a 5-fold targeted HBoV1 mAb, 15C6. Surprisingly, recognition was observed for one out of three 3-fold targeted mAbs, 12C1, indicating some structural similarity at this region. In addition, GBoV1, tested against 40 human sera, showed the similar rates of seropositivity as HBoV1. Immunogenic reactivity against parvoviral vectors is a significant barrier to efficient gene delivery. This study is a step towards optimizing bocaparvovirus vectors with antibody escape properties.
- Department of Biochemistry and Molecular Biology, Center for Structural Biology, College of Medicine, University of Florida, Gainesville, FL 32610, USA.
Organizational Affiliation: