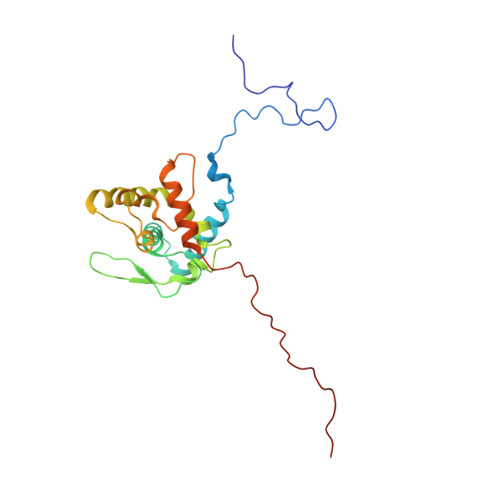
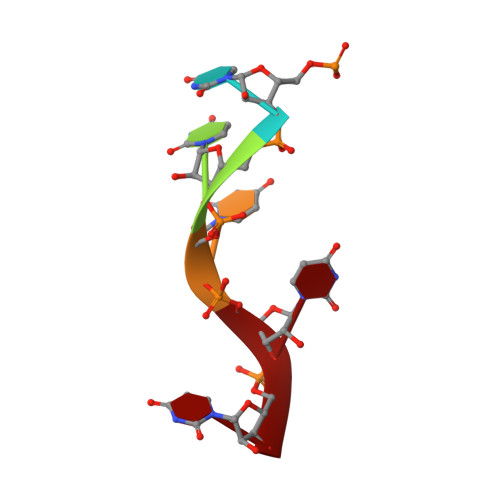
Structure of Turnip mosaic virus and its viral-like particles.
Cuesta, R., Yuste-Calvo, C., Gil-Carton, D., Sanchez, F., Ponz, F., Valle, M.(2019) Sci Rep 9: 15396-15396
- PubMed: 31659175 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-51823-4
- Primary Citation Related Structures:
6T34 - PubMed Abstract:
Turnip mosaic virus (TuMV), a potyvirus, is a flexible filamentous plant virus that displays a helical arrangement of coat protein copies (CPs) bound to the ssRNA genome. TuMV is a bona fide representative of the Potyvirus genus, one of most abundant groups of plant viruses, which displays a very wide host range. We have studied by cryoEM the structure of TuMV virions and its viral-like particles (VLPs) to explore the role of the interactions between proteins and RNA in the assembly of the virions. The results show that the CP-RNA interaction is needed for the correct orientation of the CP N-terminal arm, a region that plays as a molecular staple between CP subunits in the fully assembled virion.
- Molecular Recognition and Host-pathogen Interactions Programme, CIC bioGUNE, Bizkaia Technology Park, 48160, Derio, Spain.
Organizational Affiliation: