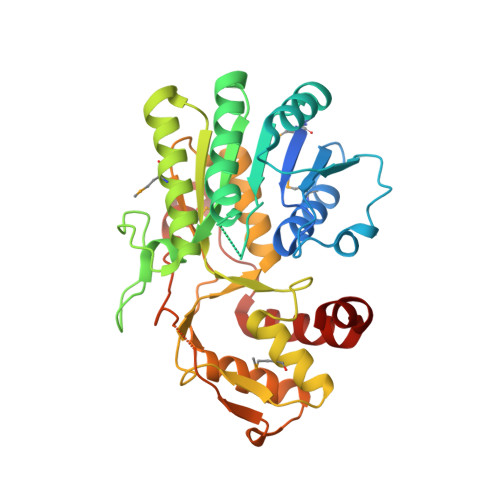
Structure of the Bacillus anthracis dTDP-l-rhamnose biosynthetic pathway enzyme: dTDP-alpha-d-glucose 4,6-dehydratase, RfbB.
Gokey, T., Halavaty, A.S., Minasov, G., Anderson, W.F., Kuhn, M.L.(2018) J Struct Biol 202: 175-181
- PubMed: 29331609 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2018.01.006
- Primary Citation Related Structures:
6BI4 - PubMed Abstract:
Many bacteria require l-rhamnose as a key cell wall component. This sugar is transferred to the cell wall using an activated donor dTDP-l-rhamnose, which is produced by the dTDP-l-rhamnose biosynthetic pathway. We determined the crystal structure of the second enzyme of this pathway dTDP-α-d-glucose 4,6-dehydratase (RfbB) from Bacillus anthracis. Interestingly, RfbB only crystallized in the presence of the third enzyme of the pathway RfbC; however, RfbC was not present in the crystal. Our work represents the first complete structural characterization of the four proteins of this pathway in a single Gram-positive bacterium.
- Department of Chemistry and Biochemistry, San Francisco State University, USA.
Organizational Affiliation: