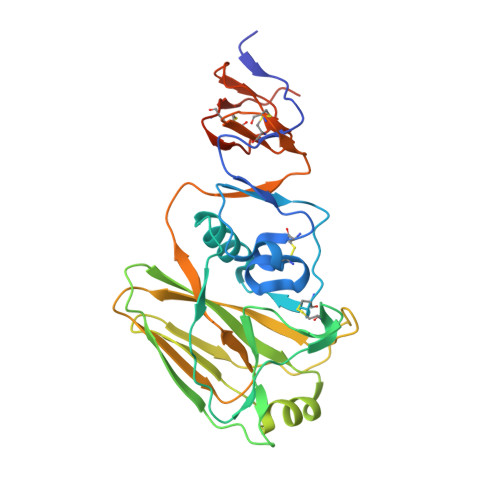
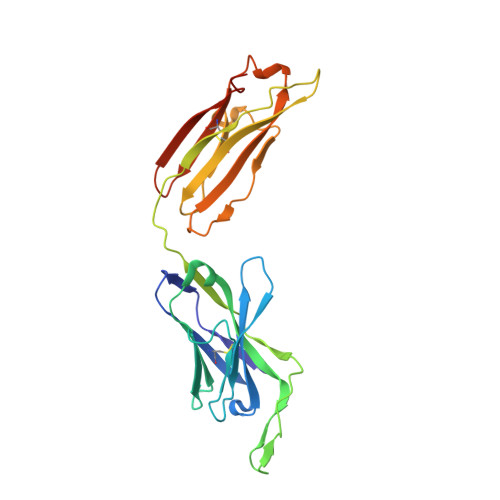
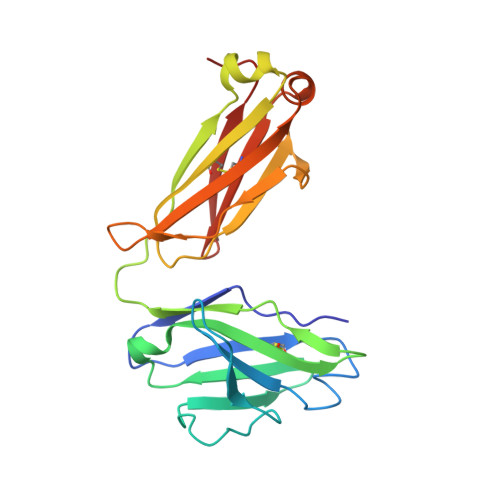
Memory B Cells that Cross-React with Group 1 and Group 2 Influenza A Viruses Are Abundant in Adult Human Repertoires.
McCarthy, K.R., Watanabe, A., Kuraoka, M., Do, K.T., McGee, C.E., Sempowski, G.D., Kepler, T.B., Schmidt, A.G., Kelsoe, G., Harrison, S.C.(2018) Immunity 48: 174-184.e9
- PubMed: 29343437 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2017.12.009
- Primary Citation Related Structures:
5W08, 5W0D - PubMed Abstract:
Human B cell antigen-receptor (BCR) repertoires reflect repeated exposures to evolving influenza viruses; new exposures update the previously generated B cell memory (Bmem) population. Despite structural similarity of hemagglutinins (HAs) from the two groups of influenza A viruses, cross-reacting antibodies (Abs) are uncommon. We analyzed Bmem compartments in three unrelated, adult donors and found frequent cross-group BCRs, both HA-head directed and non-head directed. Members of a clonal lineage from one donor had a BCR structure similar to that of a previously described Ab, encoded by different gene segments. Comparison showed that both Abs contacted the HA receptor-binding site through long heavy-chain third complementarity determining regions. Affinities of the clonal-lineage BCRs for historical influenza-virus HAs from both group 1 and group 2 viruses suggested that serial responses to seasonal influenza exposures had elicited the lineage and driven affinity maturation. We propose that appropriate immunization regimens might elicit a comparably broad response.
- Laboratory of Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: