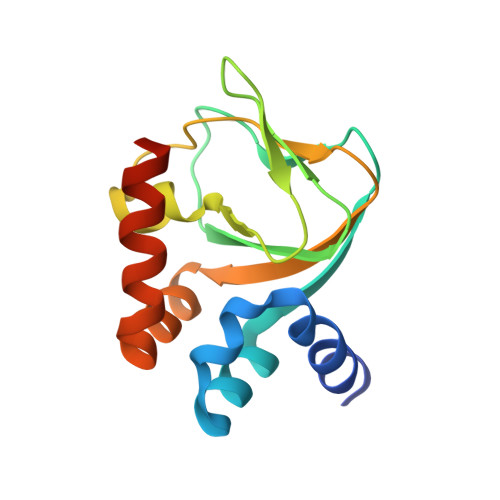
Structural and Energetic Analysis of Activation by a Cyclic Nucleotide Binding Domain.
Altieri, S.L., Clayton, G.M., Silverman, W.R., Olivares, A.O., De La Cruz, E.M., Thomas, L.R., Morais-Cabral, J.H.(2008) J Mol Biology 381: 655-669
- PubMed: 18619611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.06.011
- Primary Citation Related Structures:
3CL1, 3CLP, 3CO2 - PubMed Abstract:
MlotiK1 is a prokaryotic homolog of cyclic-nucleotide-dependent ion channels that contains an intracellular C-terminal cyclic nucleotide binding (CNB) domain. X-ray structures of the CNB domain have been solved in the absence of ligand and bound to cAMP. Both the full-length channel and CNB domain fragment are easily expressed and purified, making MlotiK1 a useful model system for dissecting activation by ligand binding. We have used X-ray crystallography to determine three new MlotiK1 CNB domain structures: a second apo configuration, a cGMP-bound structure, and a second cAMP-bound structure. In combination, the five MlotiK1 CNB domain structures provide a unique opportunity for analyzing, within a single protein, the structural differences between the apo state and the bound state, and the structural variability within each state. With this analysis as a guide, we have probed the nucleotide selectivity and importance of specific residue side chains in ligand binding and channel activation. These data help to identify ligand-protein interactions that are important for ligand dependence in MlotiK1 and, more globally, in the class of nucleotide-dependent proteins.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520-8114, USA.
Organizational Affiliation: