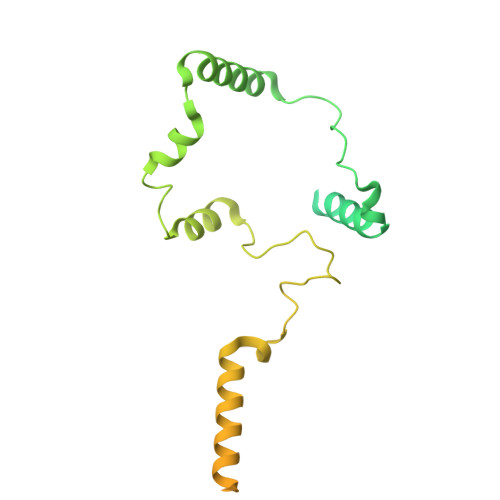
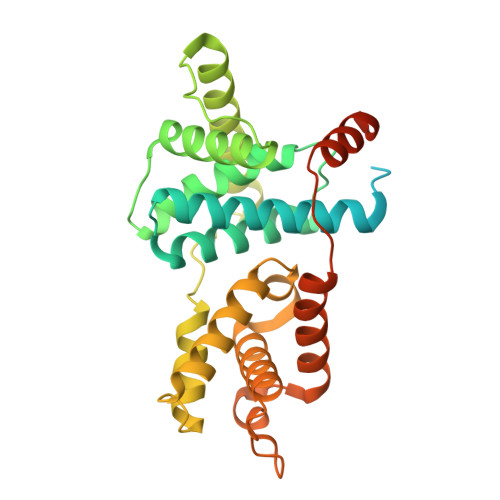
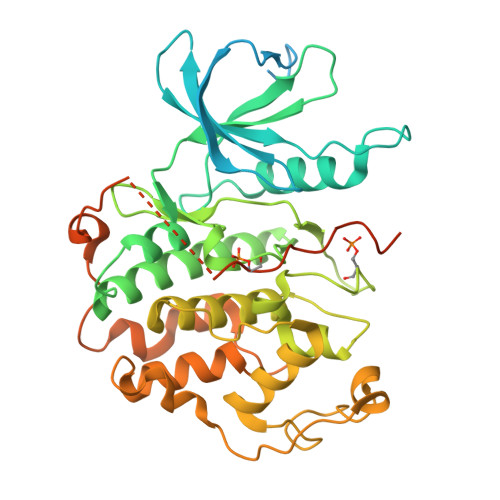
Cryo-EM structures of the CDK11-cyclin L-SAP30BP complex reveal mechanisms of CDK11 regulation.
McGeoch, A.J.S., Cushing, V.I., Roumeliotis, T.I., Cronin, N.B., Hearnshaw, S.J., Choudhary, J.S., Alfieri, C., Greber, B.J.(2026) Nat Commun
- PubMed: 42034640 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-72329-4
- Primary Citation Related Structures:
9QJJ, 9QJN, 9QKT, 9QKZ, 9QL1 - PubMed Abstract:
The cyclin-dependent kinase CDK11 functions in transcription, mitotic progression, and mRNA splicing. Specifically, spliceosome activation during the B to B act transition depends on phosphorylation of the U2 snRNP component SF3B1 by the CDK11-cyclin L-SAP30BP complex. Here, we present the structure of this spliceosome-activating CDK-cyclin complex, determined by cryogenic electron microscopy at 2.3 Å resolution. Our structure and biochemical experiments show that SAP30BP forms extensive interactions with cyclin L2, thereby stabilising it, and forms critical interactions with the C-terminal kinase lobe of CDK11 that promote complex assembly. Destabilisation of cyclin L2 in the absence of SAP30BP suggests that these principles are applicable to all CDK11-cyclin L complexes. Furthermore, we identify a pseudo-substrate sequence near the CDK11 C-terminus and provide evidence for a role of this segment in CDK11 auto-regulation. Finally, the structure of the CDK11-cyclin L-SAP30BP complex bound to the clinical high-affinity CDK11 inhibitor OTS964 and a comparison to OTS964-bound off-target complexes provide insight into the mechanism of OTS964 selectivity and specificity.
- Division of Structural Biology, The Institute of Cancer Research, London, UK.
Organizational Affiliation: