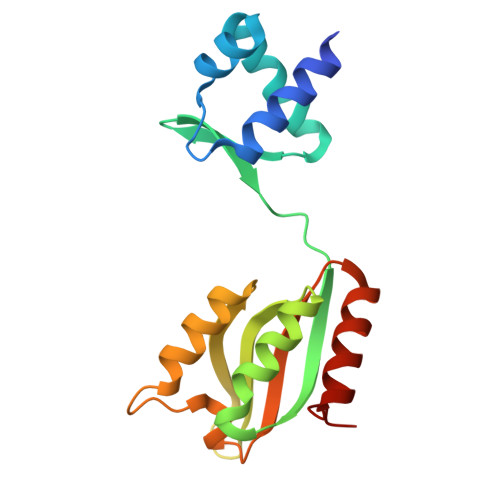
The activation of the metal-containing regulatory protein NiaR from Thermotoga maritima by its effector, nicotinic acid.
Cheng, W.C.D., Li, Y., Nakashima, M., Moenne-Loccoz, P., Rush, K.W., Glasfeld, A.(2025) J Biol Inorg Chem 30: 169-179
- PubMed: 39899144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00775-025-02096-y
- Primary Citation Related Structures:
9EBR - PubMed Abstract:
NiaR is a regulatory protein that represses the expression of proteins involved in the de novo biosynthesis and uptake of nicotinic acid (NA), with NA acting as a co-repressor. The previously published structure of NiaR from Thermotoga maritima (TmNiaR) identified it as a functional homodimer containing a transition metal ion in a suspected NA-binding pocket. Here, we present the crystal structure of NA bound to the iron-metalated form of TmNiaR. Supported by spectroscopic and solution studies, this structure shows that NA binds to a protein-bound ferrous ion via its ring nitrogen. In addition, the carboxylate group on NA interacts with Tyr108 from the dyad-related subunit, repositioning the likely DNA-binding domains of the dimer to promote high-affinity interactions with DNA operators. The specificity of TmNiaR for NA can be explained by the hydrogen bonding scheme within the NA-binding pocket.
- Department of Chemistry, Reed College, Portland, OR, 97202, USA.
Organizational Affiliation: