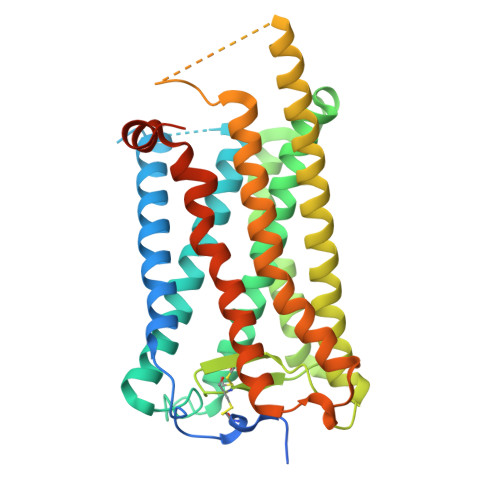
Structural insights into the mechanism of activation and inhibition of the prostaglandin D2 receptor 1.
Davoudinasab, B., Raskovalov, A., Lee, W., Kim, D., Kim, H., Lam, J.H., Han, G.W., Katritch, V., Cherezov, V.(2025) Nat Commun 16: 8944-8944
- PubMed: 41062467 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64002-z
- Primary Citation Related Structures:
9AU0, 9E9S, 9EE5, 9EI5, 9EKH - PubMed Abstract:
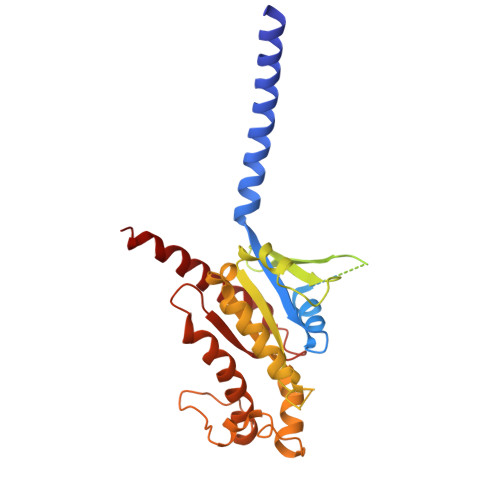
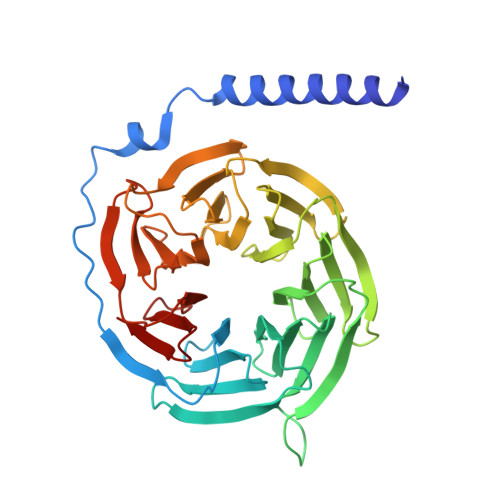
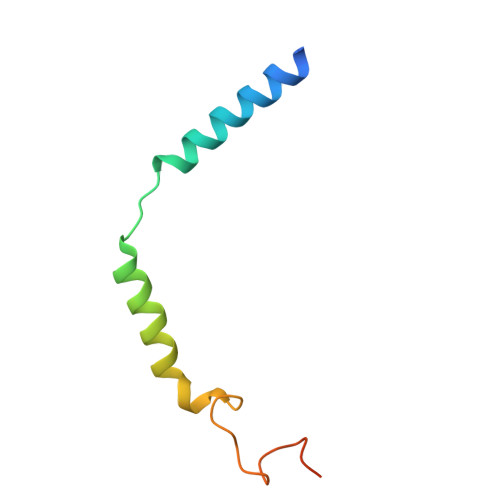
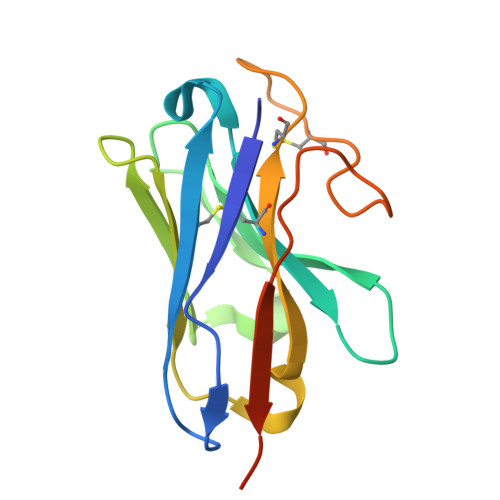
The prostaglandin D2 receptor 1 (DP1), a member of the prostanoid G protein-coupled receptor (GPCR) family, plays critical roles in allergic responses, sleep regulation, immune modulation, and vasodilation. Here, we present five high-resolution cryo-electron microscopy (cryo-EM) structures of the human DP1 receptor, including an apo structure, two inactive state structures bound to two different inverse agonists developed by ONO Pharmaceutical, and two active state structures in complex with the G s protein and bound to the endogenous agonist PGD2 and its selective derivative BW245C. Structural analysis, complemented by molecular dynamics simulations and site-directed mutagenesis, reveals key residues involved in ligand recognition and suggests a distinct activation mechanism for DP1, which lacks most of the conserved class A GPCR motifs. Notably, the unique residue K76 within the conserved sodium pocket acts as a major activation switch, while amphiphilic helix 8 adopts an unconventional orientation essential for receptor function. These findings offer valuable insights into the structure and function of prostanoid receptors and may facilitate the development of therapeutics targeting DP1.
- Department of Chemistry, University of Southern California, Los Angeles, CA, USA.
Organizational Affiliation: