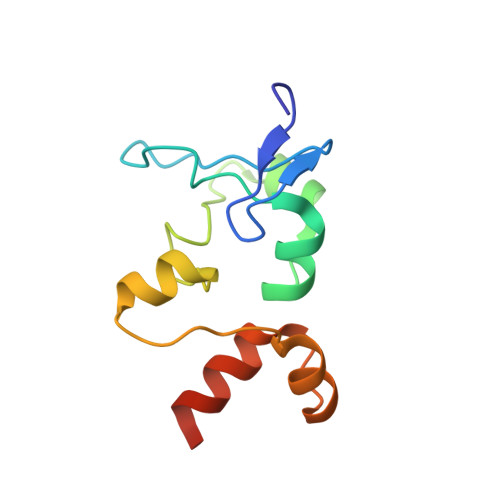
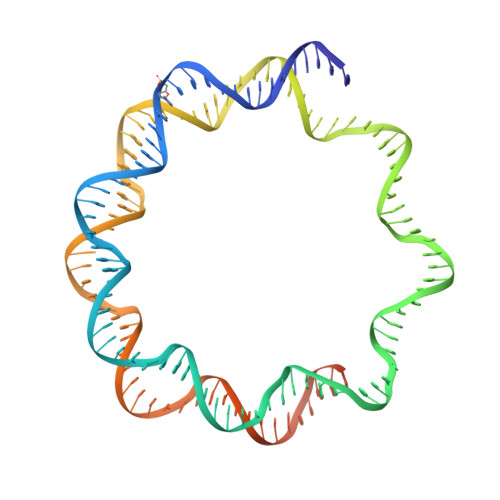
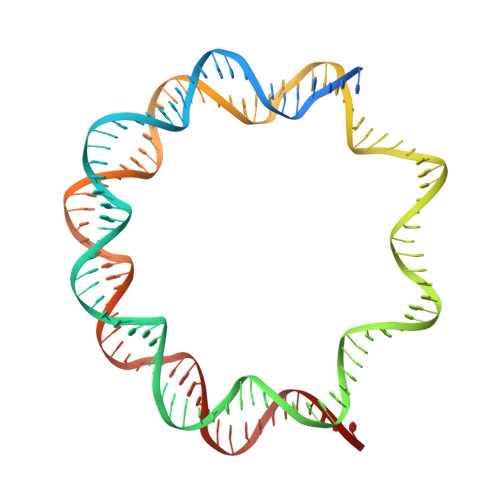
CDCA7 is an evolutionarily conserved hemimethylated DNA sensor in eukaryotes.
Wassing, I.E., Nishiyama, A., Shikimachi, R., Jia, Q., Kikuchi, A., Hiruta, M., Sugimura, K., Hong, X., Chiba, Y., Peng, J., Jenness, C., Nakanishi, M., Zhao, L., Arita, K., Funabiki, H.(2024) Sci Adv 10: eadp5753-eadp5753
- PubMed: 39178260 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adp5753
- Primary Citation Related Structures:
8YV8 - PubMed Abstract:
Mutations of the SNF2 family ATPase HELLS and its activator CDCA7 cause immunodeficiency, centromeric instability, and facial anomalies syndrome, characterized by DNA hypomethylation at heterochromatin. It remains unclear why CDCA7-HELLS is the sole nucleosome remodeling complex whose deficiency abrogates the maintenance of DNA methylation. We here identify the unique zinc-finger domain of CDCA7 as an evolutionarily conserved hemimethylation-sensing zinc finger (HMZF) domain. Cryo-electron microscopy structural analysis of the CDCA7-nucleosome complex reveals that the HMZF domain can recognize hemimethylated CpG in the outward-facing DNA major groove within the nucleosome core particle, whereas UHRF1, the critical activator of the maintenance methyltransferase DNMT1, cannot. CDCA7 recruits HELLS to hemimethylated chromatin and facilitates UHRF1-mediated H3 ubiquitylation associated with replication-uncoupled maintenance DNA methylation. We propose that the CDCA7-HELLS nucleosome remodeling complex assists the maintenance of DNA methylation on chromatin by sensing hemimethylated CpG that is otherwise inaccessible to UHRF1 and DNMT1.
- Laboratory of Chromosome and Cell Biology, The Rockefeller University, New York, NY 10065, USA.
Organizational Affiliation: