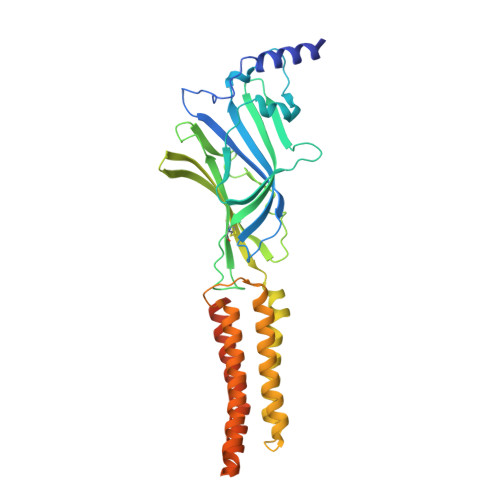
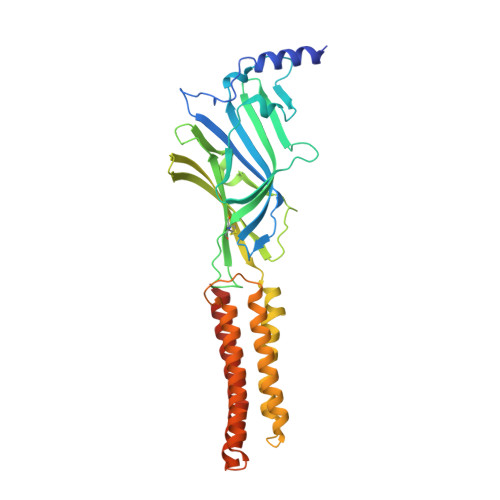
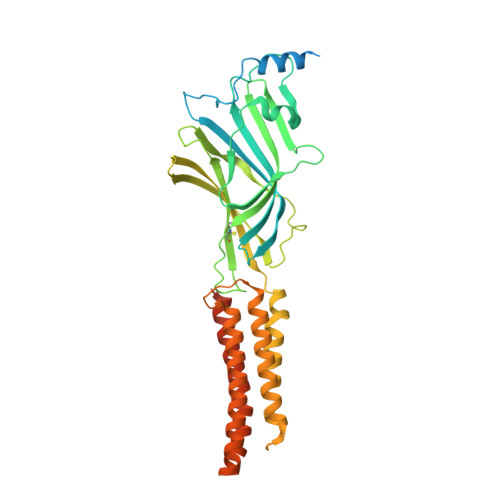
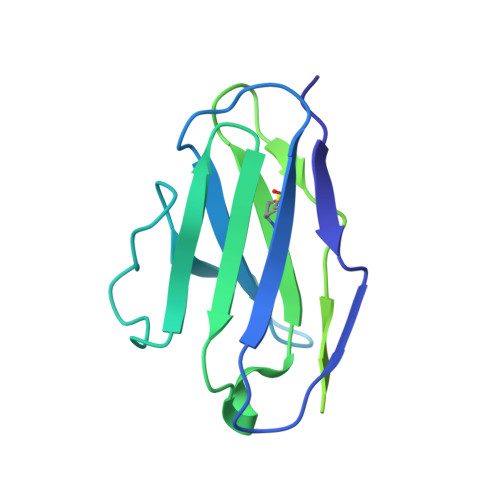
Structural insights into GABA A receptor potentiation by Quaalude.
Chojnacka, W., Teng, J., Kim, J.J., Jensen, A.A., Hibbs, R.E.(2024) Nat Commun 15: 5244-5244
- PubMed: 38898000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-49471-y
- Primary Citation Related Structures:
8VQY, 8VRN - PubMed Abstract:
Methaqualone, a quinazolinone marketed commercially as Quaalude, is a central nervous system depressant that was used clinically as a sedative-hypnotic, then became a notorious recreational drug in the 1960s-80s. Due to its high abuse potential, medical use of methaqualone was eventually prohibited, yet it persists as a globally abused substance. Methaqualone principally targets GABA A receptors, which are the major inhibitory neurotransmitter-gated ion channels in the brain. The restricted status and limited accessibility of methaqualone have contributed to its pharmacology being understudied. Here, we use cryo-EM to localize the GABA A receptor binding sites of methaqualone and its more potent derivative, PPTQ, to the same intersubunit transmembrane sites targeted by the general anesthetics propofol and etomidate. Both methaqualone and PPTQ insert more deeply into subunit interfaces than the previously-characterized modulators. Binding of quinazolinones to this site results in widening of the extracellular half of the ion-conducting pore, following a trend among positive allosteric modulators in destabilizing the hydrophobic activation gate in the pore as a mechanism for receptor potentiation. These insights shed light on the underexplored pharmacology of quinazolinones and further elucidate the molecular mechanisms of allosteric GABA A receptor modulation through transmembrane binding sites.
- Biomedical Sciences Graduate Program, University of California San Diego, La Jolla, CA, USA.
Organizational Affiliation: