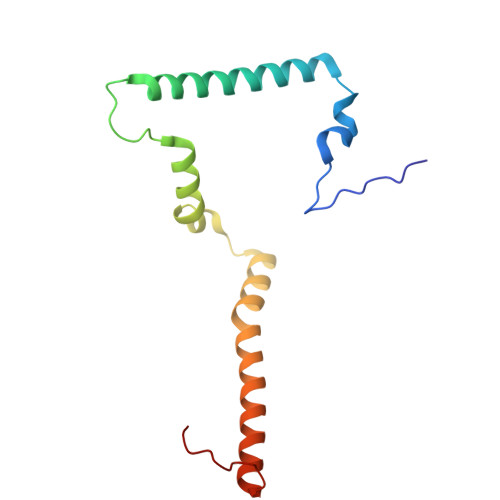
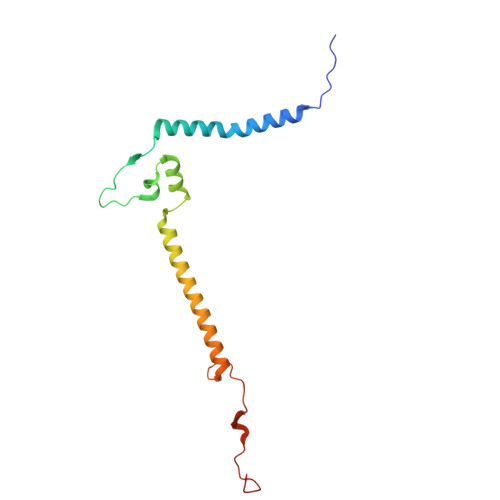
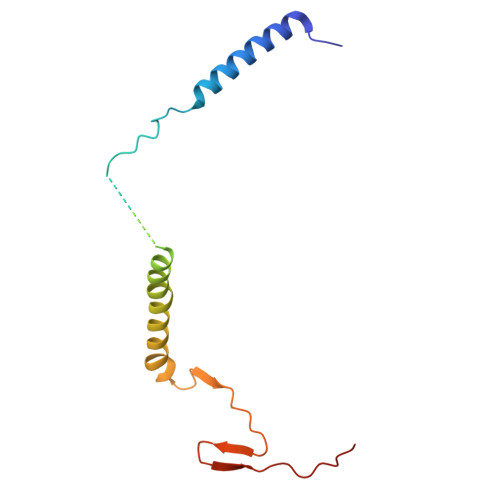
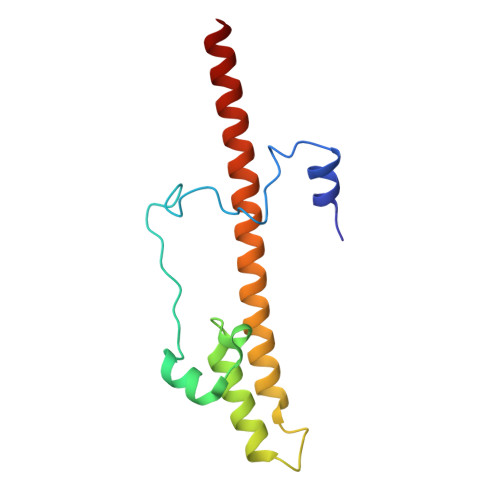
Hydrophilic metformin and hydrophobic biguanides inhibit mitochondrial complex I by distinct mechanisms.
He, Z., Teng, F., Yang, Y., Guo, R., Wu, M., Han, F., Tian, H., Wang, J., Hu, Y., Jiang, Y., Zhang, L., Xu, C., Yang, F., Zhou, J., Zhang, S., Letts, J.A., Zhou, R., Zhou, L.(2026) Nat Struct Mol Biol 33: 100-111
- PubMed: 41214295 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01710-6
- Primary Citation Related Structures:
8ZO8, 8ZOU, 8ZSK, 8ZSL, 8ZSM, 8ZSN, 8ZSO, 8ZSQ, 8ZXZ, 9J6H, 9J6W - PubMed Abstract:
Metformin is the only antihyperglycemic biguanide targeting type 2 diabetes mellitus with proven safety. Although a mechanism of action involving tight inhibition of the respiratory complex I has been proposed for hydrophobic biguanides, it remains elusive for the hydrophilic metformin, whose excellent pharmacological tolerance depends on weak complex I inhibition without competitive nature. Here we solved cryo-electron microscopy structures of the metformin-bound porcine respirasome. Our structural and kinetic data are consistent with a model in which metformin enters complex I only in its open state and becomes trapped at the ubiquinone redox site by ubiquinone-induced conformational closing of the enzyme. By contrast, the hydrophobic proguanil alone occupies both the entrance and the redox site of the ubiquinone channel in open and closed complex I and is kinetically consistent with competitive inhibition with conformation-dependent affinities. Our data provide the molecular basis for metformin's well-known superior properties, such as a wide therapeutic window and positive ubiquinone cooperativity, leading to its clinical success and facilitating future therapeutic developments.
- Department of Biophysics and Department of Critical Care Medicine of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou, China.
Organizational Affiliation: