Structural and spectroscopic insights into fucoxanthin chlorophyll a/c-binding proteins of diatoms in diverse oligomeric states.
Zhou, C., Feng, Y., Li, Z., Shen, L., Li, X., Wang, Y., Han, G., Kuang, T., Liu, C., Shen, J.R., Wang, W.(2024) Plant Commun 5: 101041-101041
- PubMed: 39030906 Search on PubMed
- DOI: https://doi.org/10.1016/j.xplc.2024.101041
- Primary Citation Related Structures:
8JP3, 8WCK, 8WCL - PubMed Abstract:
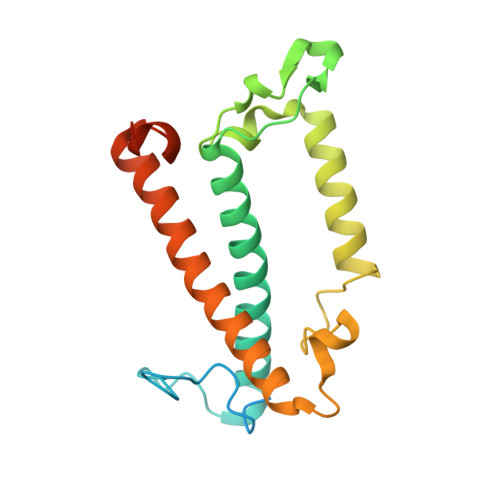
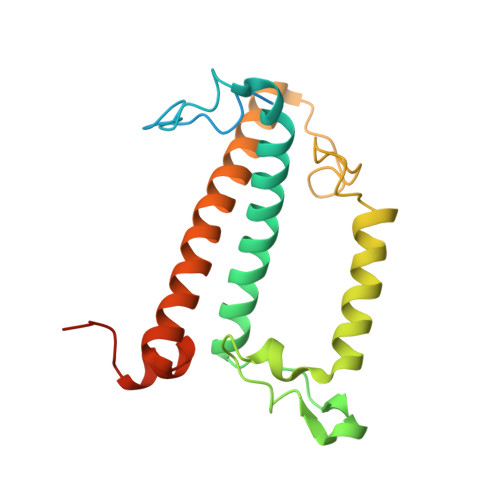
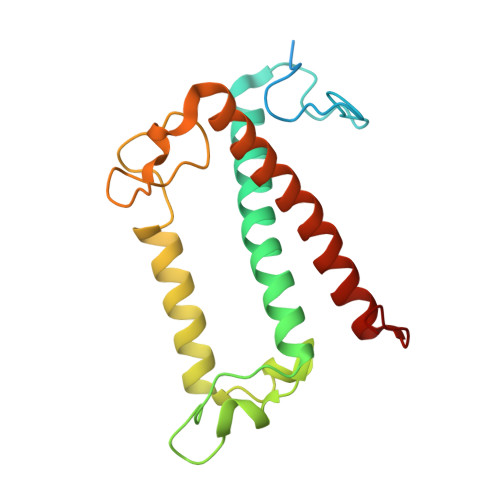
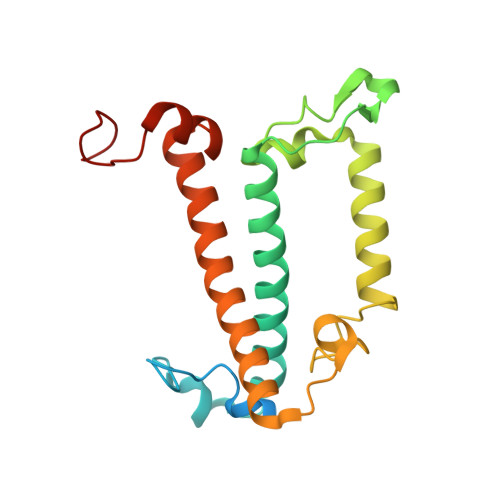
Diatoms, a group of prevalent marine algae, significantly contribute to global primary productivity. Their substantial biomass is linked to enhanced absorption of blue-green light underwater, facilitated by fucoxanthin chlorophyll a/c-binding proteins (FCPs), exhibiting oligomeric diversity across diatom species. Utilizing mild CN-PAGE analysis on solubilized thylakoid membranes, we displayed monomeric, dimeric, trimeric, tetrameric and pentameric FCPs in diatoms. Mass spectrometry analysis revealed each oligomeric FCP has specific protein compositions, constituting a large Lhcf family of FCP antennas. In addition, we resolved the structures of Thalassiosira pseudonana FCP (Tp-FCP) homotrimer and Chaetoceros gracilis FCP (Cg-FCP) pentamer by cryo-electron microscopy at 2.73 Å and 2.65 Å resolutions, respectively. The distinct pigment composition and organization in various oligomeric FCPs change their blue-green light-harvesting, excitation energy transfer pathways. In comparison to dimeric and trimeric FCPs, Cg-FCP tetramer and Cg-FCP pentamer exhibit stronger absorption by Chls c, red-shifted and broader Chl a fluorescence emission, as well as more robust circular dichroism signals originating from Chl a-carotenoid dimers. These spectroscopic characteristics indicate that Chl a molecules in Cg-FCP tetramer and Cg-FCP pentamer are more heterogeneous than in both dimers and Tp-FCP trimer. The structural and spectroscopic insights provided by this study contribute to a better understanding of the mechanisms that empower diatoms to adapt to fluctuating light environments.
- Photosynthesis Research Center, Key Laboratory of Photobiology, Institute of Botany, Chinese Academy of Sciences, Beijing 100093, P. R. China; University of Chinese Academy of Sciences, Beijing 100049, P. R. China.
Organizational Affiliation: