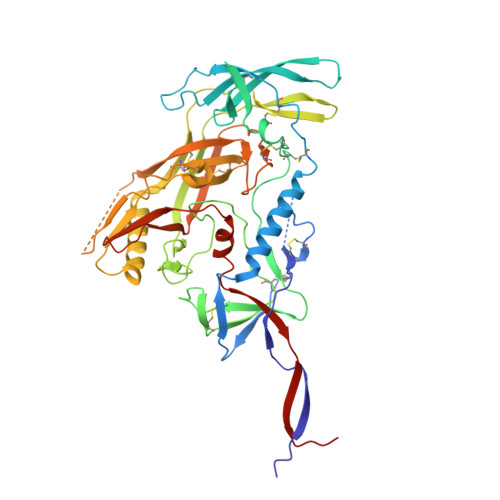
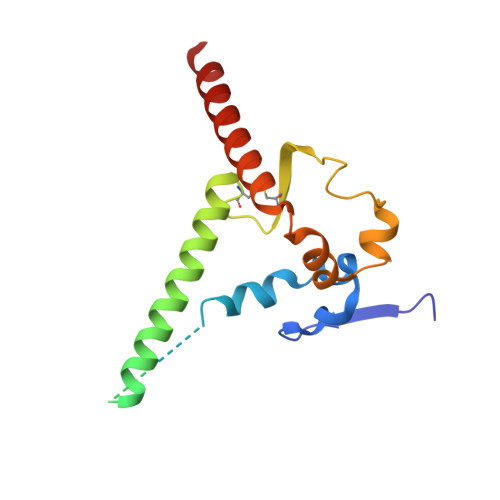
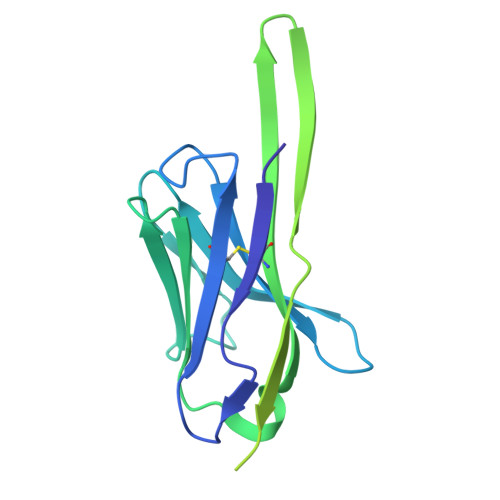
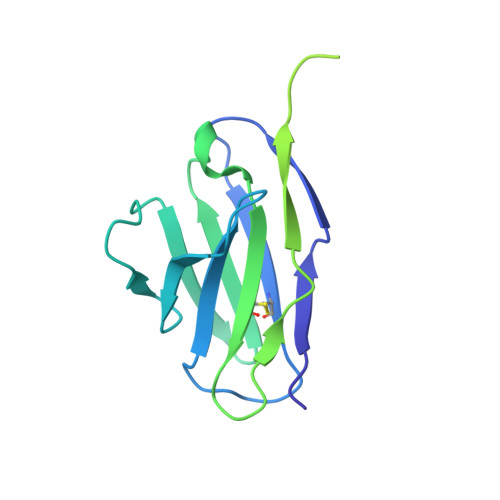
Potent and broad HIV-1 neutralization in fusion peptide-primed SHIV-infected macaques.
Wang, H., Cheng, C., Dal Santo, J.L., Shen, C.H., Bylund, T., Henry, A.R., Howe, C.A., Hwang, J., Morano, N.C., Morris, D.J., Pletnev, S., Roark, R.S., Zhou, T., Hansen, B.T., Hoyt, F.H., Johnston, T.S., Wang, S., Zhang, B., Ambrozak, D.R., Becker, J.E., Bender, M.F., Changela, A., Chaudhary, R., Corcoran, M., Corrigan, A.R., Foulds, K.E., Guo, Y., Lee, M., Li, Y., Lin, B.C., Liu, T., Louder, M.K., Mandolesi, M., Mason, R.D., McKee, K., Nair, V., O'Dell, S., Olia, A.S., Ou, L., Pegu, A., Raju, N., Rawi, R., Roberts-Torres, J., Sarfo, E.K., Sastry, M., Schaub, A.J., Schmidt, S.D., Schramm, C.A., Schwartz, C.L., Smith, S.C., Stephens, T., Stuckey, J., Teng, I.T., Todd, J.P., Tsybovsky, Y., Van Wazer, D.J., Wang, S., Doria-Rose, N.A., Fischer, E.R., Georgiev, I.S., Karlsson Hedestam, G.B., Sheng, Z., Woodward, R.A., Douek, D.C., Koup, R.A., Pierson, T.C., Shapiro, L., Shaw, G.M., Mascola, J.R., Kwong, P.D.(2024) Cell 187: 7214-7231.e23
- PubMed: 39471811 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2024.10.003
- Primary Citation Related Structures:
8TDX, 8TE7, 8TJR, 8TJS, 8TKC, 8TL2, 8TL3, 8TL4, 8TL5, 8TNU, 8TO7, 8TO9, 8TOP - PubMed Abstract:
An antibody-based HIV-1 vaccine will require the induction of potent cross-reactive HIV-1-neutralizing responses. To demonstrate feasibility toward this goal, we combined vaccination targeting the fusion-peptide site of vulnerability with infection by simian-human immunodeficiency virus (SHIV). In four macaques with vaccine-induced neutralizing responses, SHIV infection boosted plasma neutralization to 45%-77% breadth (geometric mean 50% inhibitory dilution [ID 50 ] ∼100) on a 208-strain panel. Molecular dissection of these responses by antibody isolation and cryo-electron microscopy (cryo-EM) structure determination revealed 15 of 16 antibody lineages with cross-clade neutralization to be directed toward the fusion-peptide site of vulnerability. In each macaque, isolated antibodies from memory B cells recapitulated the plasma-neutralizing response, with fusion-peptide-binding antibodies reaching breadths of 40%-60% (50% inhibitory concentration [IC 50 ] < 50 μg/mL) and total lineage-concentrations estimates of 50-200 μg/mL. Longitudinal mapping indicated that these responses arose prior to SHIV infection. Collectively, these results provide in vivo molecular examples for one to a few B cell lineages affording potent, broadly neutralizing plasma responses.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: