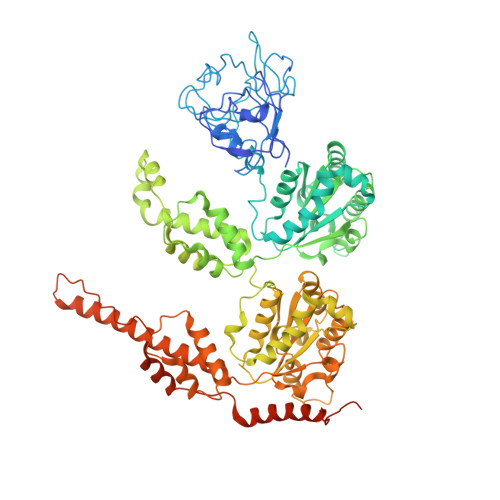
A non-symmetrical p97 conformation initiates a multistep recruitment of Ufd1/Npl4.
Arie, M., Matzov, D., Karmona, R., Szenkier, N., Stanhill, A., Navon, A.(2024) iScience 27: 110061-110061
- PubMed: 38947518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2024.110061
- Primary Citation Related Structures:
8PQX, 8R0E, 8RS9, 8RSB, 8RSC - PubMed Abstract:
In vitro experiments and cryo-EM structures of p97 and its cofactor, Ufd1/Npl4 (UN), elucidated substrate processing. Yet, the structural transitions and the related ATPase cycle upon UN binding remain unresolved. We captured two discrete conformations: One in which D1 protomers are ATP bound, while the D2 subunits are in the ADP state, presumably required for substrate engagement with the D2 pore; and a heterologous nucleotide state within the D1 ring in which only two NTDs are in the "up" ATP state that favors UN binding. Further analysis suggests that initially, UN binds p97's non-symmetrical conformation, this association promotes a structural transition upon which five NTDs shift to an "up" state and are poised to bind ATP. The UBXL domain of Npl4 was captured bound to an NTD in the ADP state, demonstrating a conformation that may provide directionality to incoming substrate and introduce the flexibility needed for substrate processing.
- Department of Immunology and Regenerative Biology, The Weizmann Institute of Science, Rehovot 7610001, Israel.
Organizational Affiliation: