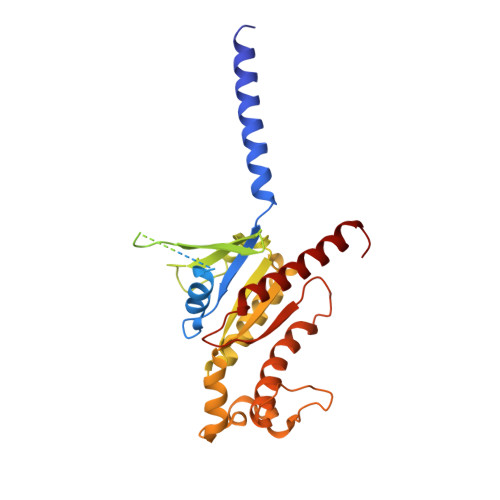
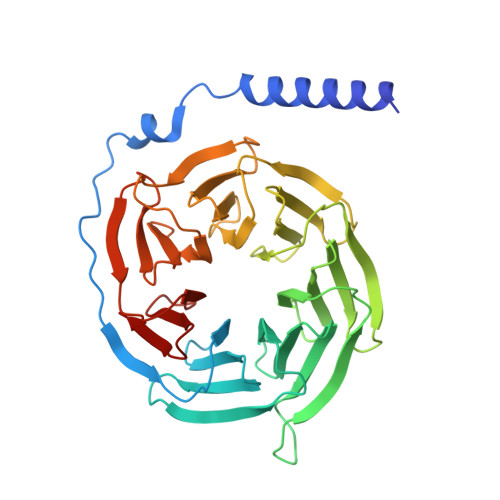
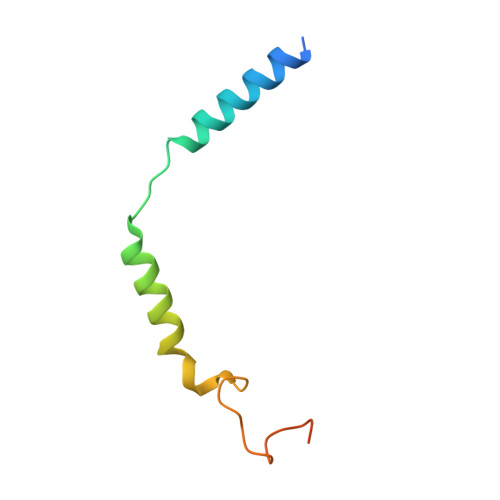
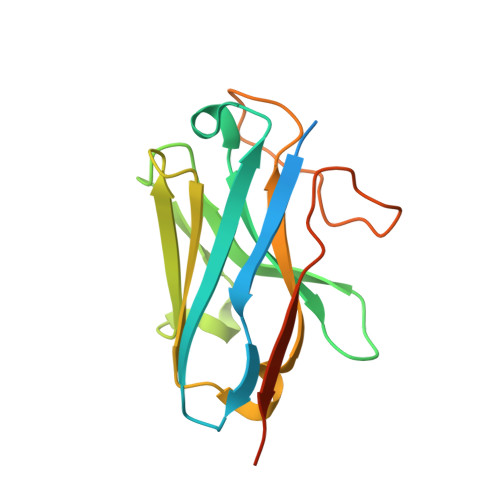
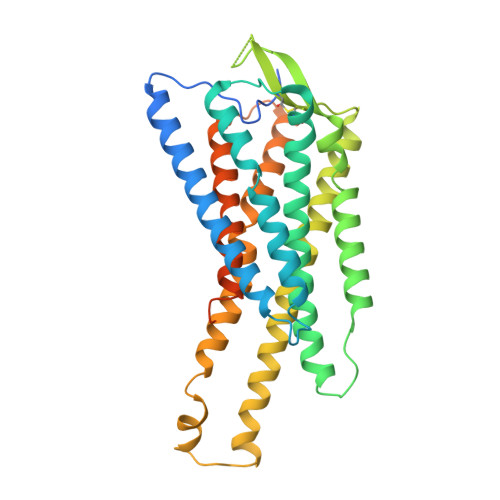
Structural basis of omega-3 fatty acid receptor FFAR4 activation and G protein coupling selectivity.
Yin, H., Inoue, A., Ma, Z., Zhu, X., Xia, R., Xu, Z., Wang, N., Duan, Y., Zhang, A., Guo, C., He, Y.(2023) Cell Res 33: 644-647
- PubMed: 37286793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-023-00835-x
- Primary Citation Related Structures:
8H4I, 8H4K, 8H4L, 8IYS - Laboratory of Receptor Structure and Signaling, HIT Center for Life Sciences, School of Life Science and Technology, Harbin Institute of Technology, Harbin, Heilongjiang, China.
Organizational Affiliation: