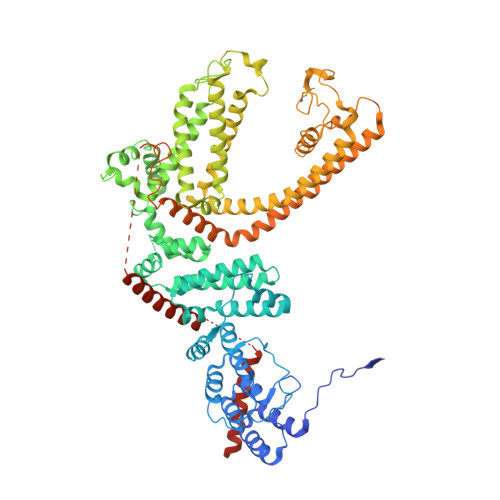
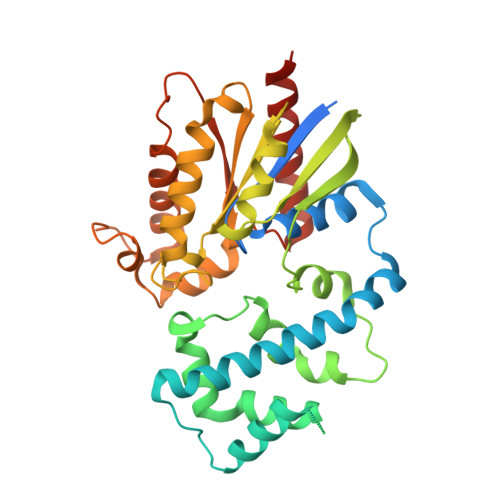
Molecular architecture of the G alpha i -bound TRPC5 ion channel.
Won, J., Kim, J., Jeong, H., Kim, J., Feng, S., Jeong, B., Kwak, M., Ko, J., Im, W., So, I., Lee, H.H.(2023) Nat Commun 14: 2550-2550
- PubMed: 37137991 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38281-3
- Primary Citation Related Structures:
7X6C, 7X6I, 8GVW, 8GVX - PubMed Abstract:
G-protein coupled receptors (GPCRs) and ion channels serve as key molecular switches through which extracellular stimuli are transformed into intracellular effects, and it has long been postulated that ion channels are direct effector molecules of the alpha subunit of G-proteins (Gα). However, no complete structural evidence supporting the direct interaction between Gα and ion channels is available. Here, we present the cryo-electron microscopy structures of the human transient receptor potential canonical 5 (TRPC5)-Gα i3 complexes with a 4:4 stoichiometry in lipid nanodiscs. Remarkably, Gα i3 binds to the ankyrin repeat edge of TRPC5 ~ 50 Å away from the cell membrane. Electrophysiological analysis shows that Gα i3 increases the sensitivity of TRPC5 to phosphatidylinositol 4,5-bisphosphate (PIP 2 ), thereby rendering TRPC5 more easily opened in the cell membrane, where the concentration of PIP 2 is physiologically regulated. Our results demonstrate that ion channels are one of the direct effector molecules of Gα proteins triggered by GPCR activation-providing a structural framework for unraveling the crosstalk between two major classes of transmembrane proteins: GPCRs and ion channels.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul, 08826, Republic of Korea.
Organizational Affiliation: