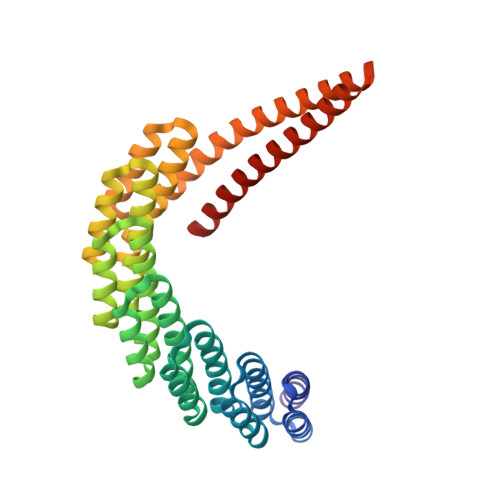
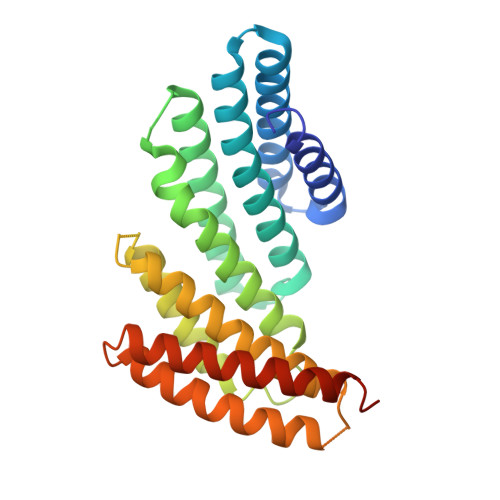
De novo design of proteins housing excitonically coupled chlorophyll special pairs.
Ennist, N.M., Wang, S., Kennedy, M.A., Curti, M., Sutherland, G.A., Vasilev, C., Redler, R.L., Maffeis, V., Shareef, S., Sica, A.V., Hua, A.S., Deshmukh, A.P., Moyer, A.P., Hicks, D.R., Swartz, A.Z., Cacho, R.A., Novy, N., Bera, A.K., Kang, A., Sankaran, B., Johnson, M.P., Phadkule, A., Reppert, M., Ekiert, D., Bhabha, G., Stewart, L., Caram, J.R., Stoddard, B.L., Romero, E., Hunter, C.N., Baker, D.(2024) Nat Chem Biol 20: 906-915
- PubMed: 38831036 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-024-01626-0
- Primary Citation Related Structures:
7UNH, 7UNI, 7UNJ, 8EVM, 8GLT - PubMed Abstract:
Natural photosystems couple light harvesting to charge separation using a 'special pair' of chlorophyll molecules that accepts excitation energy from the antenna and initiates an electron-transfer cascade. To investigate the photophysics of special pairs independently of the complexities of native photosynthetic proteins, and as a first step toward creating synthetic photosystems for new energy conversion technologies, we designed C 2 -symmetric proteins that hold two chlorophyll molecules in closely juxtaposed arrangements. X-ray crystallography confirmed that one designed protein binds two chlorophylls in the same orientation as native special pairs, whereas a second designed protein positions them in a previously unseen geometry. Spectroscopy revealed that the chlorophylls are excitonically coupled, and fluorescence lifetime imaging demonstrated energy transfer. The cryo-electron microscopy structure of a designed 24-chlorophyll octahedral nanocage with a special pair on each edge closely matched the design model. The results suggest that the de novo design of artificial photosynthetic systems is within reach of current computational methods.
- Institute for Protein Design, University of Washington, Seattle, WA, USA. ennist@uw.edu.
Organizational Affiliation: