DNA-encoded library-enabled discovery of proximity-inducing small molecules.
Mason, J.W., Chow, Y.T., Hudson, L., Tutter, A., Michaud, G., Westphal, M.V., Shu, W., Ma, X., Tan, Z.Y., Coley, C.W., Clemons, P.A., Bonazzi, S., Berst, F., Briner, K., Liu, S., Zecri, F.J., Schreiber, S.L.(2024) Nat Chem Biol 20: 170-179
- PubMed: 37919549 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01458-4
- Primary Citation Related Structures:
8EWV - PubMed Abstract:
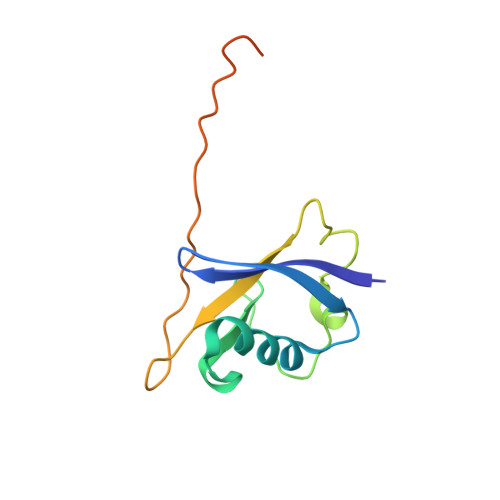
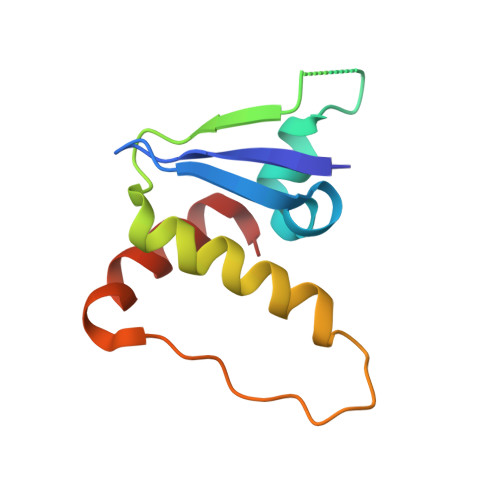
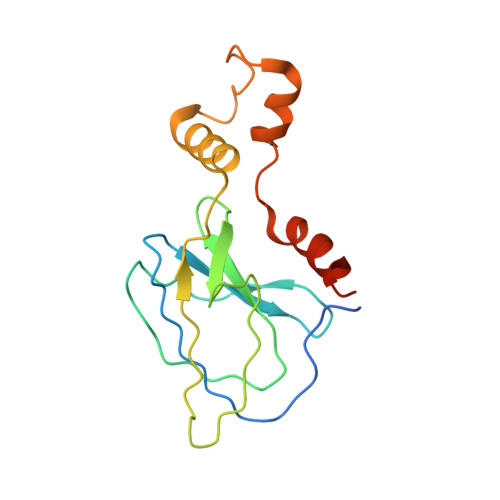
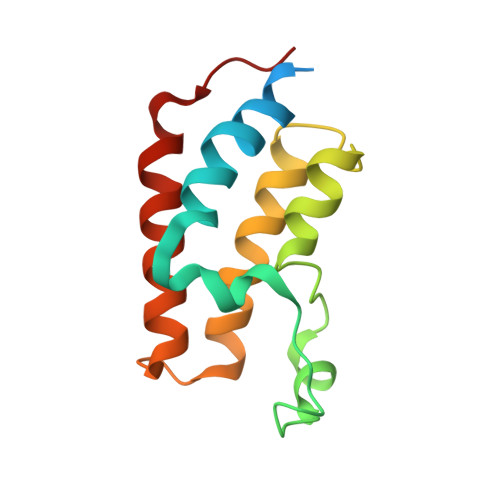
Small molecules that induce protein-protein associations represent powerful tools to modulate cell circuitry. We sought to develop a platform for the direct discovery of compounds able to induce association of any two preselected proteins, using the E3 ligase von Hippel-Lindau (VHL) and bromodomains as test systems. Leveraging the screening power of DNA-encoded libraries (DELs), we synthesized ~1 million DNA-encoded compounds that possess a VHL-targeting ligand, a variety of connectors and a diversity element generated by split-and-pool combinatorial chemistry. By screening our DEL against bromodomains in the presence and absence of VHL, we could identify VHL-bound molecules that simultaneously bind bromodomains. For highly barcode-enriched library members, ternary complex formation leading to bromodomain degradation was confirmed in cells. Furthermore, a ternary complex crystal structure was obtained for our most enriched library member with BRD4 BD1 and a VHL complex. Our work provides a foundation for adapting DEL screening to the discovery of proximity-inducing small molecules.
- Chemical Biology and Therapeutics Science, Broad Institute, Cambridge, MA, USA.
Organizational Affiliation: