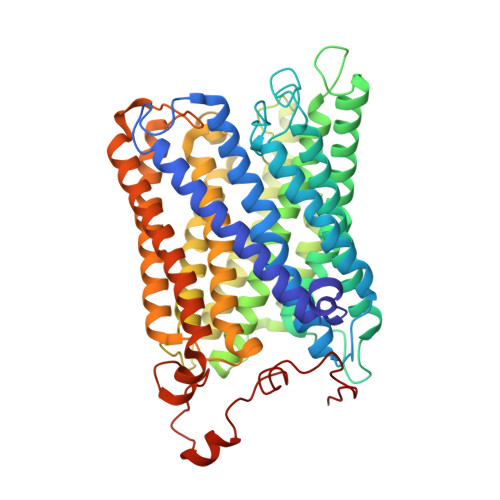
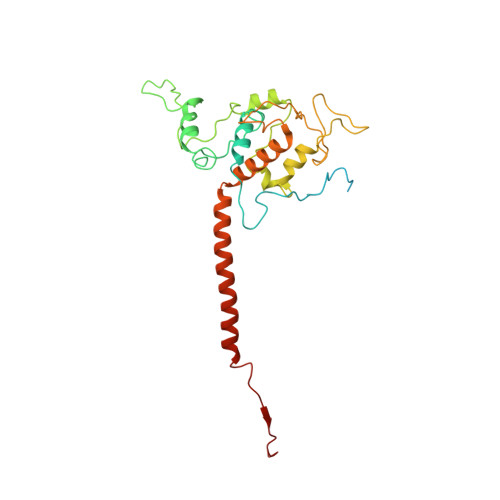
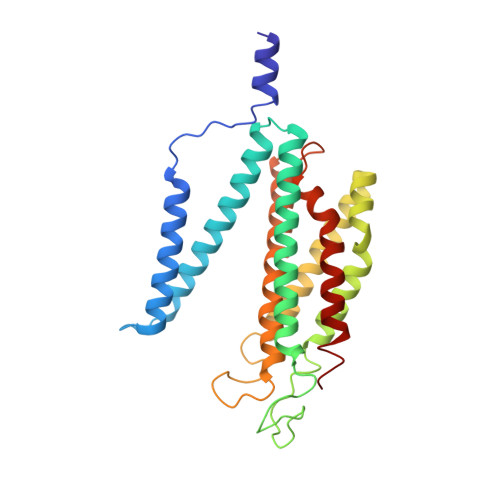
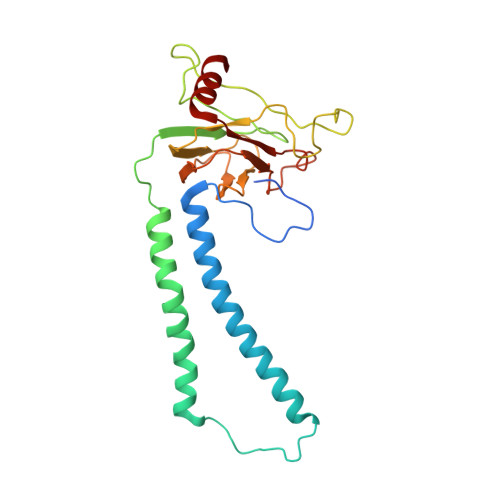
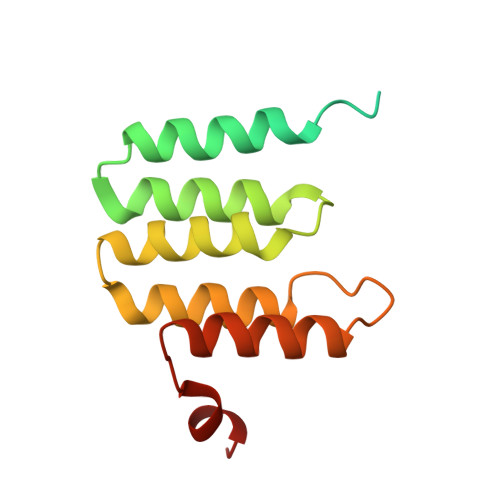
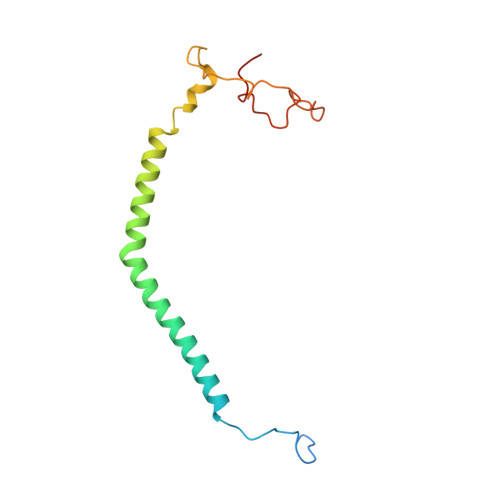
Structural insights into cardiolipin replacement by phosphatidylglycerol in a cardiolipin-lacking yeast respiratory supercomplex.
Hryc, C.F., Mallampalli, V.K.P.S., Bovshik, E.I., Azinas, S., Fan, G., Serysheva, I.I., Sparagna, G.C., Baker, M.L., Mileykovskaya, E., Dowhan, W.(2023) Nat Commun 14: 2783-2783
- PubMed: 37188665 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-38441-5
- Primary Citation Related Structures:
8E7S, 8EC0 - PubMed Abstract:
Cardiolipin is a hallmark phospholipid of mitochondrial membranes. Despite established significance of cardiolipin in supporting respiratory supercomplex organization, a mechanistic understanding of this lipid-protein interaction is still lacking. To address the essential role of cardiolipin in supercomplex organization, we report cryo-EM structures of a wild type supercomplex (IV 1 III 2 IV 1 ) and a supercomplex (III 2 IV 1 ) isolated from a cardiolipin-lacking Saccharomyces cerevisiae mutant at 3.2-Å and 3.3-Å resolution, respectively, and demonstrate that phosphatidylglycerol in III 2 IV 1 occupies similar positions as cardiolipin in IV 1 III 2 IV 1 . Lipid-protein interactions within these complexes differ, which conceivably underlies the reduced level of IV 1 III 2 IV 1 and high levels of III 2 IV 1 and free III 2 and IV in mutant mitochondria. Here we show that anionic phospholipids interact with positive amino acids and appear to nucleate a phospholipid domain at the interface between the individual complexes, which dampen charge repulsion and further stabilize interaction, respectively, between individual complexes.
- Department of Biochemistry and Molecular Biology, McGovern Medical School at the University of Texas Health Science Center, Houston, Texas, USA.
Organizational Affiliation: