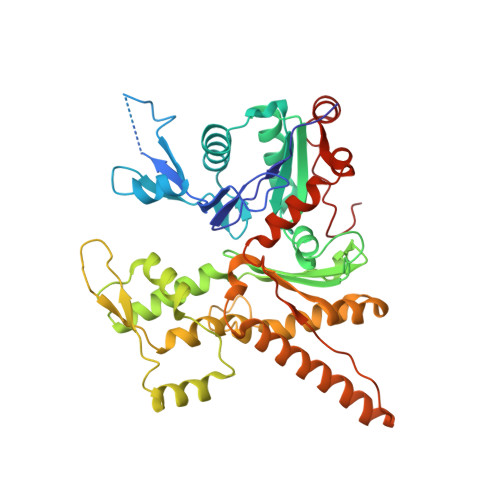
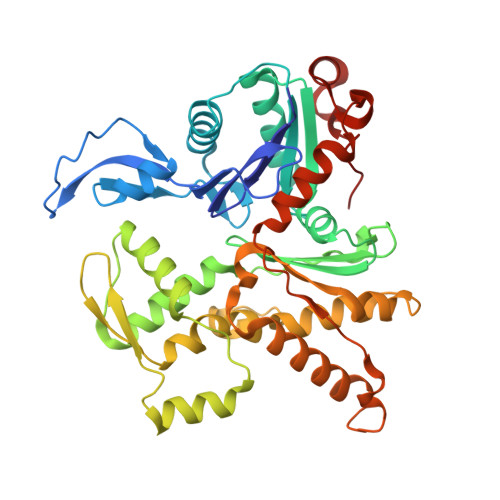
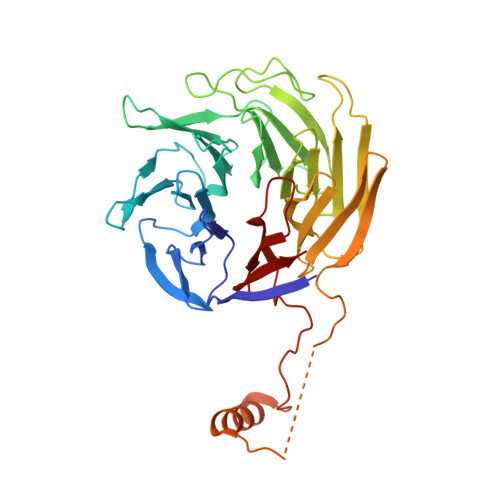
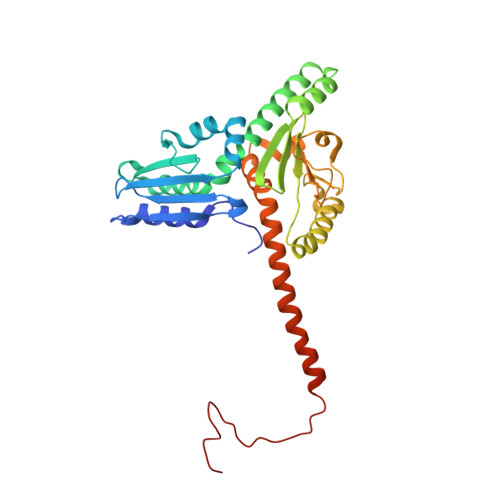
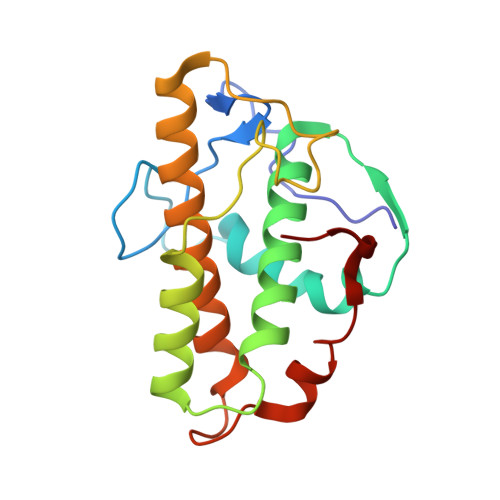
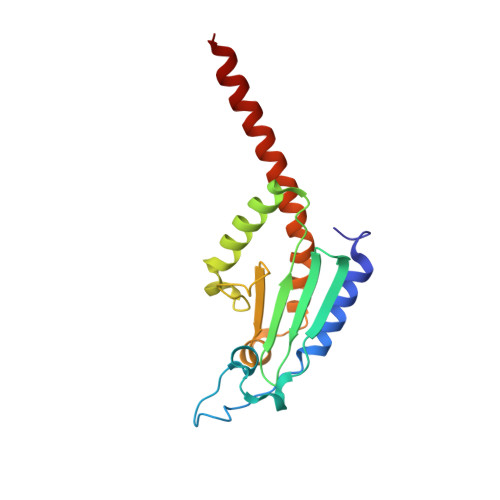
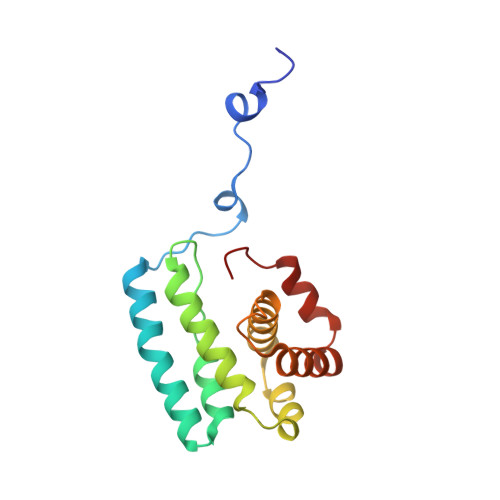
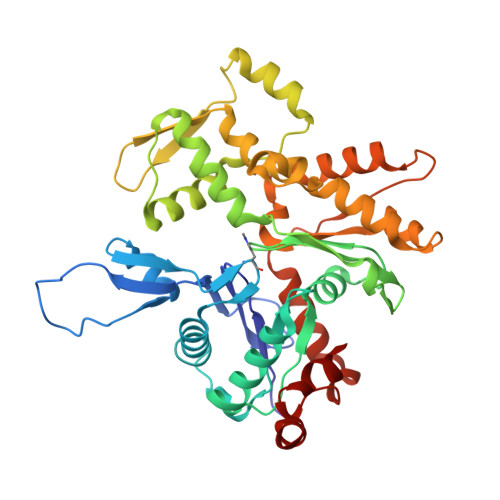
Mechanism of actin filament branch formation by Arp2/3 complex revealed by a high-resolution cryo-EM structureof the branch junction.
Chou, S.Z., Chatterjee, M., Pollard, T.D.(2022) Proc Natl Acad Sci U S A 119: e2206722119-e2206722119
- PubMed: 36442092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2206722119
- Primary Citation Related Structures:
8E9B - PubMed Abstract:
We reconstructed the structure of actin filament branch junctions formed by fission yeast Arp2/3 complex at 3.5 Å resolution from images collected by electron cryo-microscopy. During specimen preparation, all of the actin subunits and Arp3 hydrolyzed their bound adenosine triphosphate (ATP) and dissociated the γ-phosphate, but Arp2 retained the γ-phosphate. Binding tightly to the side of the mother filament and nucleating the daughter filament growing as a branch requires Arp2/3 complex to undergo a dramatic conformational change where two blocks of structure rotate relative to each other about 25° to align Arp2 and Arp3 as the first two subunits in the branch. During branch formation, Arp2/3 complex acquires more than 8,000 Å 2 of new buried surface, accounting for the stability of the branch. Inactive Arp2/3 complex binds only transiently to the side of an actin filament, because its conformation allows only a subset of the interactions found in the branch junction.
- Department of Molecular Cellular and Developmental Biology, Yale University, New Haven, CT 06520.
Organizational Affiliation: