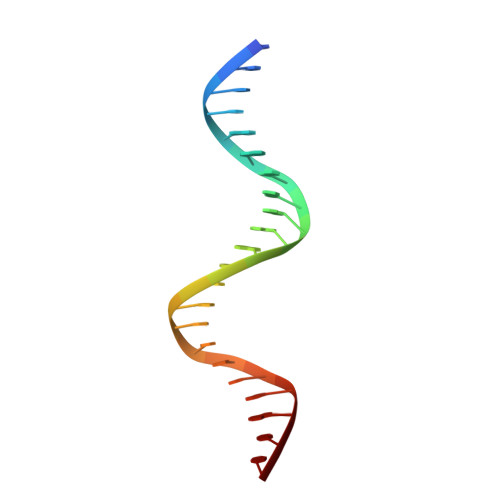
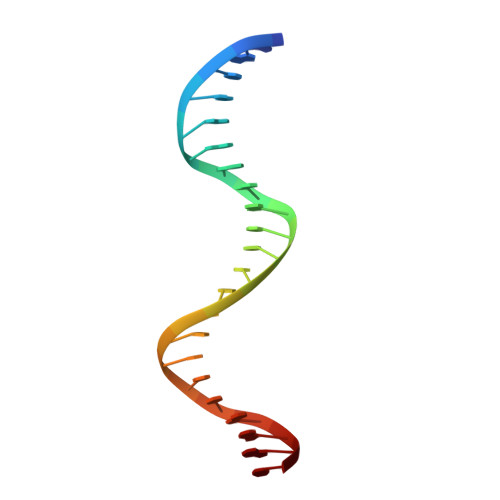
Structural basis for transcription factor ZBTB7A recognition of DNA and effects of ZBTB7A somatic mutations that occur in human acute myeloid leukemia.
Ren, R., Horton, J.R., Chen, Q., Yang, J., Liu, B., Huang, Y., Blumenthal, R.M., Zhang, X., Cheng, X.(2023) J Biological Chem 299: 102885-102885
- PubMed: 36626981 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.102885
- Primary Citation Related Structures:
7N5U, 7N5V, 7N5W, 8E3D, 8E3E - PubMed Abstract:
ZBTB7A belongs to a small family of transcription factors having three members in humans (7A, 7B, and 7C). They share a BTB/POZ protein interaction domain at the amino end and a zinc-finger DNA-binding domain at the carboxyl end. They control the transcription of a wide range of genes, having varied functions in hematopoiesis, oncogenesis, and metabolism (in particular glycolysis). ZBTB7A-binding profiles at gene promoters contain a consensus G(a/c)CCC motif, followed by a CCCC sequence in some instances. Structural and mutational investigations suggest that DNA-specific contacts with the four-finger tandem array of ZBTB7A are formed sequentially, initiated from ZF1-ZF2 binding to G(a/c)CCC before spreading to ZF3-ZF4, which bind the DNA backbone and the 3' CCCC sequence, respectively. Here, we studied some mutations found in t(8;21)-positive acute myeloid leukemia patients that occur within the ZBTB7A DNA-binding domain. We determined that these mutations generally impair ZBTB7A DNA binding, with the most severe disruptions resulting from mutations in ZF1 and ZF2, and the least from a frameshift mutation in ZF3 that results in partial mislocalization. Information provided here on ZBTB7A-DNA interactions is likely applicable to ZBTB7B/C, which have overlapping functions with ZBTB7A in controlling primary metabolism.
- Department of Epigenetics and Molecular Carcinogenesis, University of Texas MD Anderson Cancer Center, Houston, Texas, USA.
Organizational Affiliation: