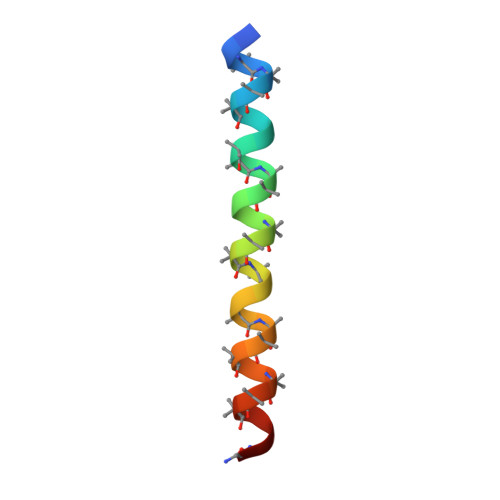
De novo design of discrete, stable 3 10 -helix peptide assemblies.
Kumar, P., Paterson, N.G., Clayden, J., Woolfson, D.N.(2022) Nature 607: 387-392
- PubMed: 35732733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-04868-x
- Primary Citation Related Structures:
7QDI, 7QDJ, 7QDK, 8BFD, 8BFE - PubMed Abstract:
The α-helix is pre-eminent in structural biology 1 and widely exploited in protein folding 2 , design 3 and engineering 4 . Although other helical peptide conformations do exist near to the α-helical region of conformational space-namely, 3 10 -helices and π-helices 5 -these occur much less frequently in protein structures. Less favourable internal energies and reduced tendencies to pack into higher-order structures mean that 3 10 -helices rarely exceed six residues in length in natural proteins, and that they tend not to form normal supersecondary, tertiary or quaternary interactions. Here we show that despite their absence in nature, synthetic peptide assemblies can be built from 3 10 -helices. We report the rational design, solution-phase characterization and an X-ray crystal structure for water-soluble bundles of 3 10 -helices with consolidated hydrophobic cores. The design uses six-residue repeats informed by analysing 3 10 -helical conformations in known protein structures, and incorporates α-aminoisobutyric acid residues. Design iterations reveal a tipping point between α-helical and 3 10 -helical folding, and identify features required for stabilizing assemblies of 3 10 -helices. This work provides principles and rules to open opportunities for designing into this hitherto unexplored region of protein-structure space.
- School of Chemistry, University of Bristol, Bristol, UK. prasun.kumar@bristol.ac.uk.
Organizational Affiliation: