Inhibition of Urease by Hydroquinones: A Structural and Kinetic Study.
Mazzei, L., Cianci, M., Ciurli, S.(2022) Chemistry 28: e202201770-e202201770
- PubMed: 35994380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/chem.202201770
- Primary Citation Related Structures:
8A18 - PubMed Abstract:
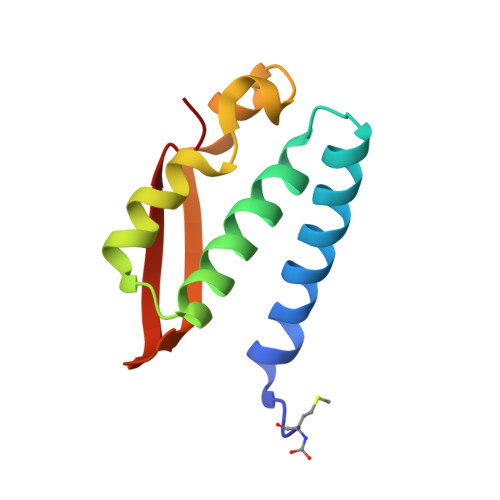
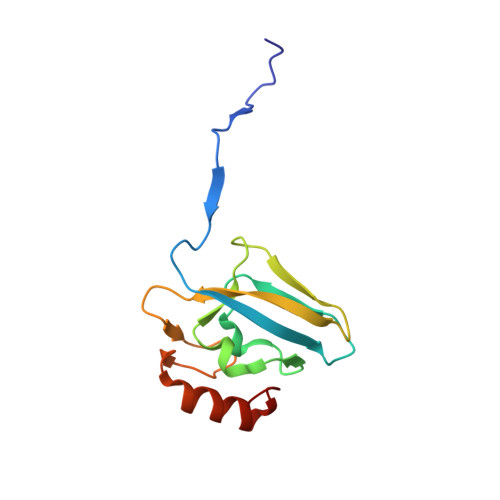
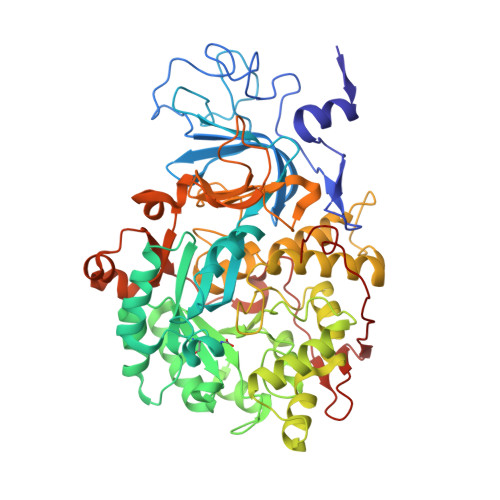
Hydroquinones are a class of organic compounds abundant in nature that result from the full reduction of the corresponding quinones. Quinones are known to efficiently inhibit urease, a Ni II -containing enzyme that catalyzes the hydrolysis of urea to yield ammonia and carbonate and acts as a virulence factor of several human pathogens, in addition to decreasing the efficiency of soil organic nitrogen fertilization. Here, we report the molecular characterization of the inhibition of urease from Sporosarcina pasteurii (SPU) and Canavalia ensiformis (jack bean, JBU) by 1,4-hydroquinone (HQ) and its methyl and tert-butyl derivatives. The 1.63-Å resolution X-ray crystal structure of the SPU-HQ complex discloses that HQ covalently binds to the thiol group of αCys322, a key residue located on a mobile protein flap directly involved in the catalytic mechanism. Inhibition kinetic data obtained for the three compounds on JBU reveals the occurrence of an irreversible inactivation process that involves a radical-based autocatalytic mechanism.
- Laboratory of Bioinorganic Chemistry Department of Pharmacy and Biotechnology (FaBiT), University of Bologna, Viale Giuseppe Fanin 40, 40127, Bologna, Italy.
Organizational Affiliation: