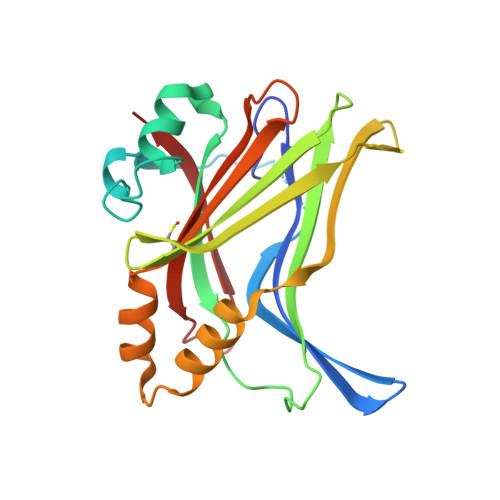
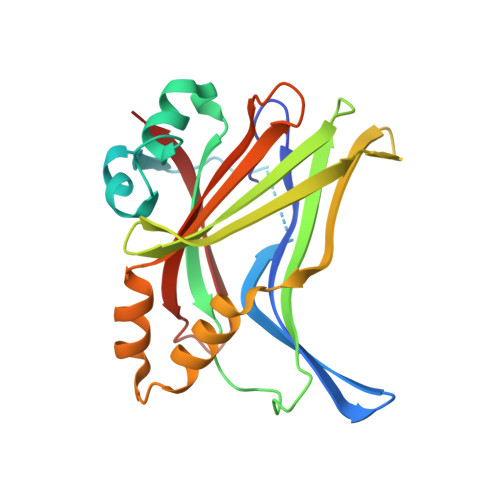
Optimization of TEAD P-Site Binding Fragment Hit into In Vivo Active Lead MSC-4106 .
Heinrich, T., Peterson, C., Schneider, R., Garg, S., Schwarz, D., Gunera, J., Seshire, A., Kotzner, L., Schlesiger, S., Musil, D., Schilke, H., Doerfel, B., Diehl, P., Bopple, P., Lemos, A.R., Sousa, P.M.F., Freire, F., Bandeiras, T.M., Carswell, E., Pearson, N., Sirohi, S., Hooker, M., Trivier, E., Broome, R., Balsiger, A., Crowden, A., Dillon, C., Wienke, D.(2022) J Med Chem 65: 9206-9229
- PubMed: 35763499 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.2c00403
- Primary Citation Related Structures:
7ZJP, 7ZJQ - PubMed Abstract:
The dysregulated Hippo pathway and, consequently, hyperactivity of the transcriptional YAP/TAZ-TEAD complexes is associated with diseases such as cancer. Prevention of YAP/TAZ-TEAD triggered gene transcription is an attractive strategy for therapeutic intervention. The deeply buried and conserved lipidation pocket (P-site) of the TEAD transcription factors is druggable. The discovery and optimization of a P-site binding fragment ( 1 ) are described. Utilizing structure-based design, enhancement in target potency was engineered into the hit, capitalizing on the established X-ray structure of TEAD1. The efforts culminated in the optimized in vivo tool MSC-4106 , which exhibited desirable potency, mouse pharmacokinetic properties, and in vivo efficacy. In close correlation to compound exposure, the time- and dose-dependent downregulation of a proximal biomarker could be shown.
- Merck Healthcare KGaA, Frankfurter Str. 250, 64293 Darmstadt, Germany.
Organizational Affiliation: