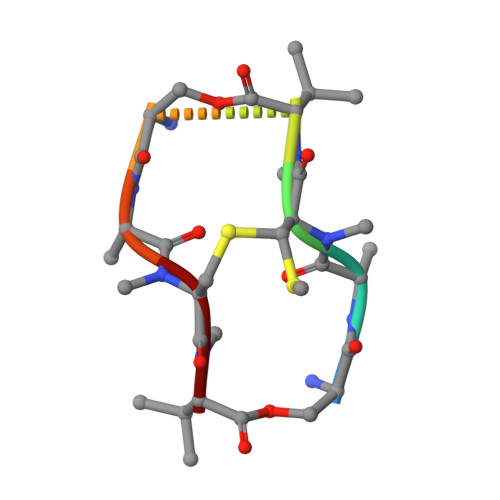
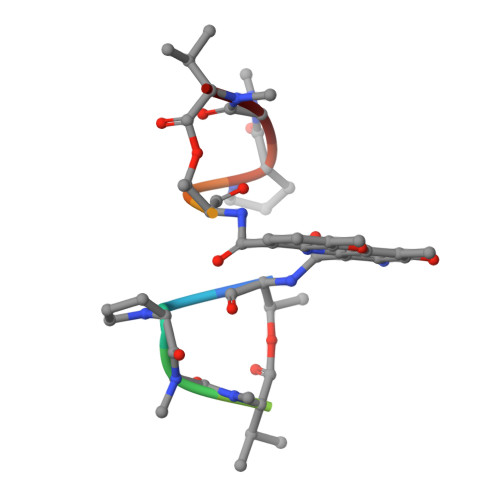
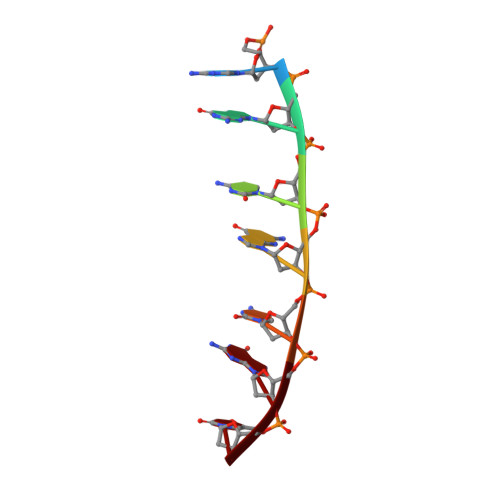
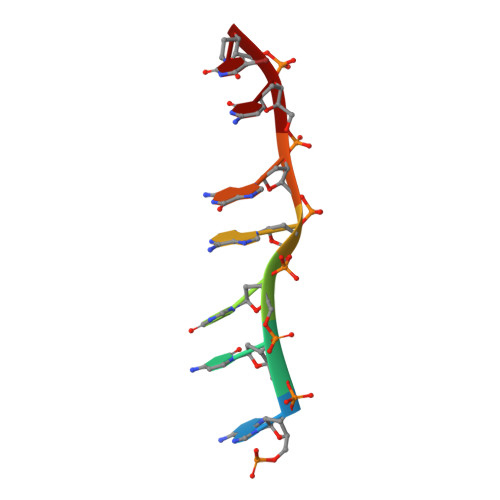
Staggered intercalation of DNA duplexes with base-pair modulation by two distinct drug molecules induces asymmetric backbone twisting and structure polymorphism.
Satange, R., Kao, S.H., Chien, C.M., Chou, S.H., Lin, C.C., Neidle, S., Hou, M.H.(2022) Nucleic Acids Res 50: 8867-8881
- PubMed: 35871296 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac629
- Primary Citation Related Structures:
7X6R, 7X97, 7X9F, 7XDJ - PubMed Abstract:
The use of multiple drugs simultaneously targeting DNA is a promising strategy in cancer therapy for potentially overcoming single drug resistance. In support of this concept, we report that a combination of actinomycin D (ActD) and echinomycin (Echi), can interact in novel ways with native and mismatched DNA sequences, distinct from the structural effects produced by either drug alone. Changes in the former with GpC and CpG steps separated by a A:G or G:A mismatch or in a native DNA with canonical G:C and C:G base pairs, result in significant asymmetric backbone twists through staggered intercalation and base pair modulations. A wobble or Watson-Crick base pair at the two drug-binding interfaces can result in a single-stranded 'chair-shaped' DNA duplex with a straight helical axis. However, a novel sugar-edged hydrogen bonding geometry in the G:A mismatch leads to a 'curved-shaped' duplex. Two non-canonical G:C Hoogsteen base pairings produce a sharply kinked duplex in different forms and a four-way junction-like superstructure, respectively. Therefore, single base pair modulations on the two drug-binding interfaces could significantly affect global DNA structure. These structures thus provide a rationale for atypical DNA recognition via multiple DNA intercalators and a structural basis for the drugs' potential synergetic use.
- Institute of Genomics and Bioinformatics, National Chung Hsing University, Taichung 402, Taiwan.
Organizational Affiliation: