Brassinosteroid-induced gene repression requires specific and tight promoter binding of BIL1/BZR1 via DNA shape readout.
Nosaki, S., Mitsuda, N., Sakamoto, S., Kusubayashi, K., Yamagami, A., Xu, Y., Bui, T.B.C., Terada, T., Miura, K., Nakano, T., Tanokura, M., Miyakawa, T.(2022) Nat Plants 8: 1440-1452
- PubMed: 36522451 Search on PubMed
- DOI: https://doi.org/10.1038/s41477-022-01289-6
- Primary Citation Related Structures:
7VN2, 7VN3, 7VN4, 7VN5, 7VN6, 7VN7, 7VN8 - PubMed Abstract:
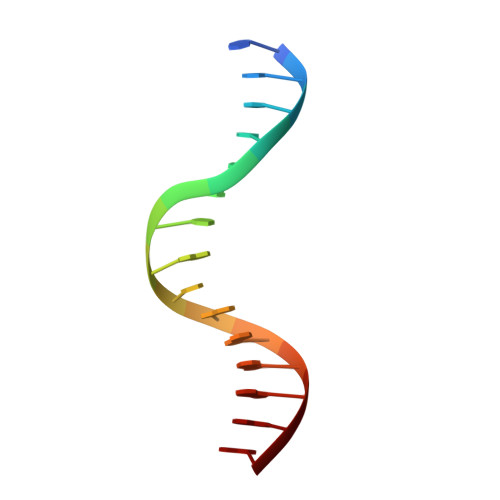
BRZ-INSENSITIVE-LONG 1 (BIL1)/BRASSINAZOLE-RESISTANT 1 (BZR1) and its homologues are plant-specific transcription factors that convert the signalling of the phytohormones brassinosteroids (BRs) to transcriptional responses, thus controlling various physiological processes in plants. Although BIL1/BZR1 upregulates some BR-responsive genes and downregulates others, the molecular mechanism underlying the dual roles of BIL1/BZR1 is still poorly understood. Here we show that BR-responsive transcriptional repression by BIL1/BZR1 requires the tight binding of BIL1/BZR1 alone to the 10 bp elements of DNA fragments containing the known 6 bp core-binding motifs at the centre. Furthermore, biochemical and structural evidence demonstrates that the selectivity for two nucleobases flanking the core motifs is realized by the DNA shape readout of BIL1/BZR1 without direct recognition of the nucleobases. These results elucidate the molecular and structural basis of transcriptional repression by BIL1/BZR1 and contribute to further understanding of the dual roles of BIL1/BZR1 in BR-responsive gene regulation.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: