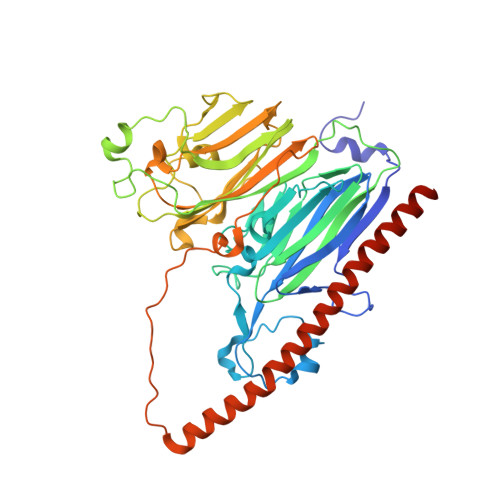
Identification of difructose dianhydride I synthase/hydrolase from an oral bacterium establishes a novel glycoside hydrolase family.
Kashima, T., Okumura, K., Ishiwata, A., Kaieda, M., Terada, T., Arakawa, T., Yamada, C., Shimizu, K., Tanaka, K., Kitaoka, M., Ito, Y., Fujita, K., Fushinobu, S.(2021) J Biological Chem 297: 101324-101324
- PubMed: 34688653 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.101324
- Primary Citation Related Structures:
7V1V, 7V1W, 7V1X - PubMed Abstract:
Fructooligosaccharides and their anhydrides are widely used as health-promoting foods and prebiotics. Various enzymes acting on β-D-fructofuranosyl linkages of natural fructan polymers have been used to produce functional compounds. However, enzymes that hydrolyze and form α-D-fructofuranosyl linkages have been less studied. Here, we identified the BBDE_2040 gene product from Bifidobacterium dentium (α-D-fructofuranosidase and difructose dianhydride I synthase/hydrolase from Bifidobacterium dentium [αFFase1]) as an enzyme with α-D-fructofuranosidase and α-D-arabinofuranosidase activities and an anomer-retaining manner. αFFase1 is not homologous with any known enzymes, suggesting that it is a member of a novel glycoside hydrolase family. When caramelized fructose sugar was incubated with αFFase1, conversions of β-D-Frup-(2→1)-α-D-Fruf to α-D-Fruf-1,2':2,1'-β-D-Frup (diheterolevulosan II) and β-D-Fruf-(2→1)-α-D-Fruf (inulobiose) to α-D-Fruf-1,2':2,1'-β-D-Fruf (difructose dianhydride I [DFA I]) were observed. The reaction equilibrium between inulobiose and DFA I was biased toward the latter (1:9) to promote the intramolecular dehydrating condensation reaction. Thus, we named this enzyme DFA I synthase/hydrolase. The crystal structures of αFFase1 in complex with β-D-Fruf and β-D-Araf were determined at the resolutions of up to 1.76 Å. Modeling of a DFA I molecule in the active site and mutational analysis also identified critical residues for catalysis and substrate binding. The hexameric structure of αFFase1 revealed the connection of the catalytic pocket to a large internal cavity via a channel. Molecular dynamics analysis implied stable binding of DFA I and inulobiose to the active site with surrounding water molecules. Taken together, these results establish DFA I synthase/hydrolase as a member of a new glycoside hydrolase family (GH172).
- Department of Biotechnology, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: