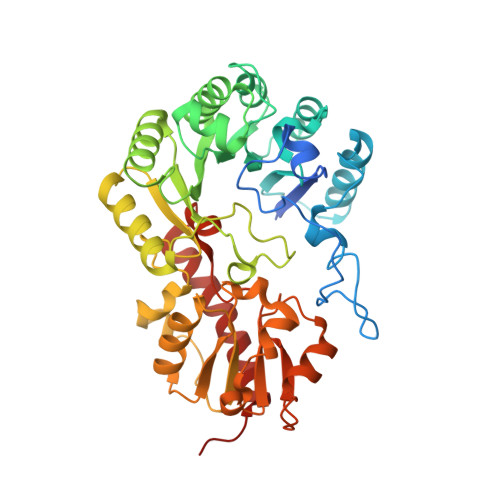
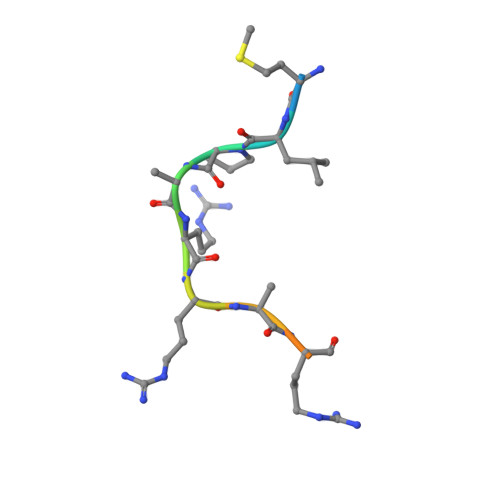
Crystallographic snapshots of a B 12 -dependent radical SAM methyltransferase.
Fyfe, C.D., Bernardo-Garcia, N., Fradale, L., Grimaldi, S., Guillot, A., Brewee, C., Chavas, L.M.G., Legrand, P., Benjdia, A., Berteau, O.(2022) Nature 602: 336-342
- PubMed: 35110733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-04355-9
- Primary Citation Related Structures:
7QBS, 7QBT, 7QBU, 7QBV - PubMed Abstract:
By catalysing the microbial formation of methane, methyl-coenzyme M reductase has a central role in the global levels of this greenhouse gas 1,2 . The activity of methyl-coenzyme M reductase is profoundly affected by several unique post-translational modifications 3-6 , such as a unique C-methylation reaction catalysed by methanogenesis marker protein 10 (Mmp10), a radical S-adenosyl-L-methionine (SAM) enzyme 7,8 . Here we report the spectroscopic investigation and atomic resolution structure of Mmp10 from Methanosarcina acetivorans, a unique B 12 (cobalamin)-dependent radical SAM enzyme 9 . The structure of Mmp10 reveals a unique enzyme architecture with four metallic centres and critical structural features involved in the control of catalysis. In addition, the structure of the enzyme-substrate complex offers a glimpse into a B 12 -dependent radical SAM enzyme in a precatalytic state. By combining electron paramagnetic resonance spectroscopy, structural biology and biochemistry, our study illuminates the mechanism by which the emerging superfamily of B 12 -dependent radical SAM enzymes catalyse chemically challenging alkylation reactions and identifies distinctive active site rearrangements to provide a structural rationale for the dual use of the SAM cofactor for radical and nucleophilic chemistry.
- Université Paris-Saclay, INRAE, AgroParisTech, Micalis Institute, ChemSyBio, Jouy-en-Josas, France.
Organizational Affiliation: