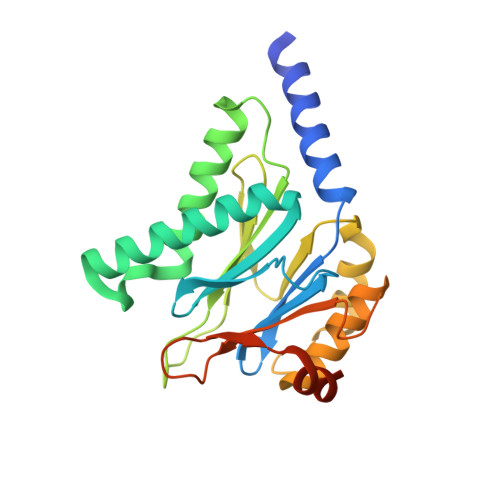
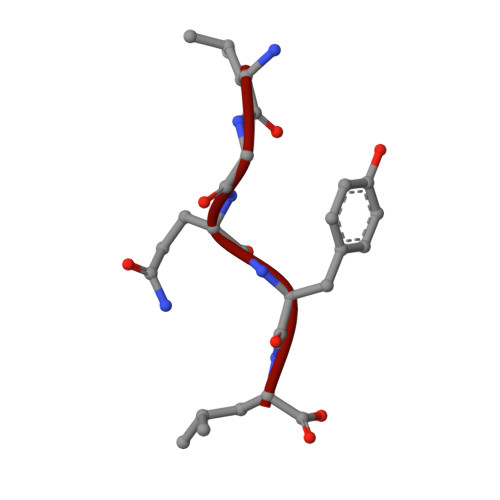
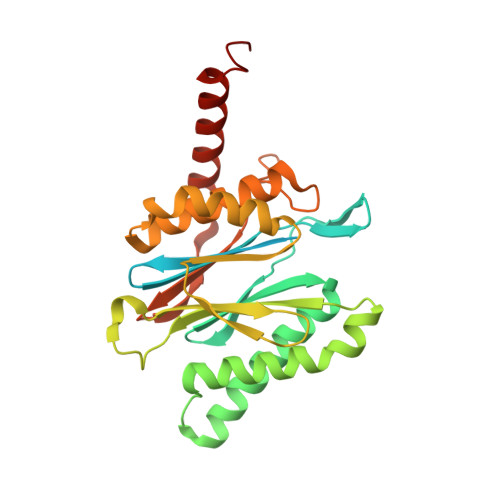
Structural basis of prokaryotic ubiquitin-like protein engagement and translocation by the mycobacterial Mpa-proteasome complex.
Kavalchuk, M., Jomaa, A., Muller, A.U., Weber-Ban, E.(2022) Nat Commun 13: 276-276
- PubMed: 35022401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27787-3
- Primary Citation Related Structures:
7PX9, 7PXA, 7PXB, 7PXC, 7PXD - PubMed Abstract:
Proteasomes are present in eukaryotes, archaea and Actinobacteria, including the human pathogen Mycobacterium tuberculosis, where proteasomal degradation supports persistence inside the host. In mycobacteria and other members of Actinobacteria, prokaryotic ubiquitin-like protein (Pup) serves as a degradation tag post-translationally conjugated to target proteins for their recruitment to the mycobacterial proteasome ATPase (Mpa). Here, we use single-particle cryo-electron microscopy to determine the structure of Mpa in complex with the 20S core particle at an early stage of pupylated substrate recruitment, shedding light on the mechanism of substrate translocation. Two conformational states of Mpa show how substrate is translocated stepwise towards the degradation chamber of the proteasome core particle. We also demonstrate, in vitro and in vivo, the importance of a structural feature in Mpa that allows formation of alternating charge-complementary interactions with the proteasome resulting in radial, rail-guided movements during the ATPase conformational cycle.
- ETH Zurich, Institute of Molecular Biology & Biophysics, CH-8093, Zurich, Switzerland.
Organizational Affiliation: