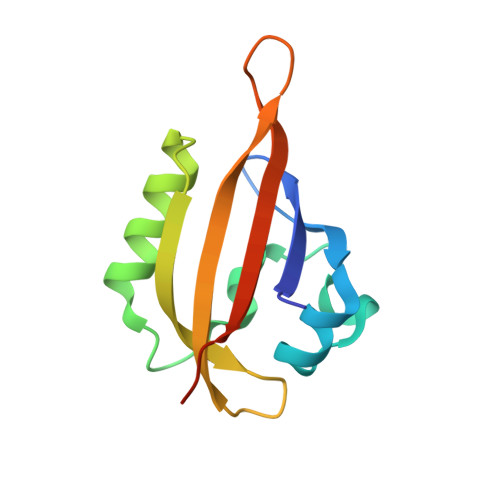
High-resolution structure of a naturally red-shifted LOV domain.
Goncharov, I.M., Smolentseva, A., Semenov, O., Natarov, I., Nazarenko, V.V., Yudenko, A., Remeeva, A., Gushchin, I.(2021) Biochem Biophys Res Commun 567: 143-147
- PubMed: 34153684 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2021.06.046
- Primary Citation Related Structures:
7OO9 - PubMed Abstract:
LOV domains are widespread photosensory modules that have also found applications in fluorescence microscopy, optogenetics, and light-driven generation of reactive oxygen species. Many of these applications require stable proteins with altered spectra. Here, we report a flavin-based fluorescent protein CisFbFP derived from Chloroflexus islandicus LOV domain-containing protein. We show that CisFbFP is thermostable, and its absorption and fluorescence spectra are red-shifted for ∼6 nm, which has not been observed for other cysteine-substituted natural LOV domains. We also provide a crystallographic structure of CisFbFP at the resolution of 1.2 Å that reveals alterations in the active site due to replacement of conservative asparagine with a serine. Finally, we discuss the possible effects of presence of cis-proline in the Aβ-Bβ loop on the protein's structure and stability. The findings provide the basis for engineering and color tuning of LOV-based tools for molecular biology.
- Research Center for Molecular Mechanisms of Aging and Age-Related Diseases, Moscow Institute of Physics and Technology, 141700, Dolgoprudny, Russia.
Organizational Affiliation: