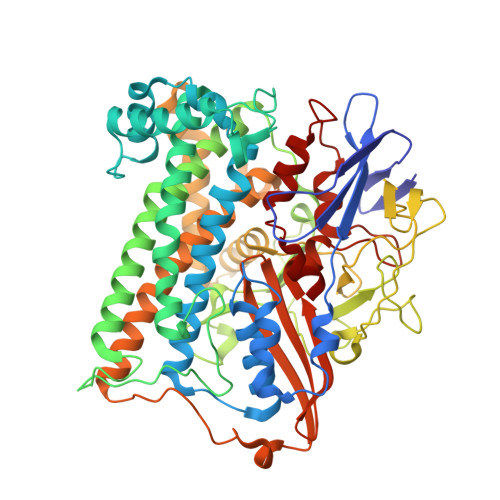
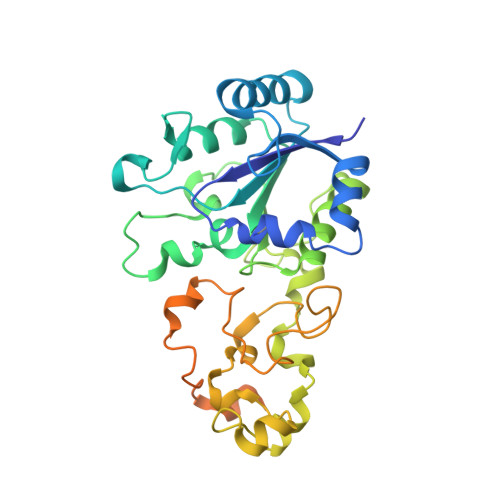
Resonance Raman spectroscopic analysis of the iron-sulfur cluster redox chain of the Ralstonia eutropha membrane-bound [NiFe]-hydrogenase
Siebert, E., Schmidt, A., Frielingsdorf, S., Kalms, J., Kuhlmann, U., Lenz, O., Scheerer, P., Zebger, I., Hildebrandt, P.(2021) J Raman Spectrosc