Crystal structure of an octameric RuvA-Holliday junction complex
Roe, S.M., Pearl, L.H.(1998) Molecular Cell 2: 361-372
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
This entry reflects an alternative modeling of the original data in: 1bvs
(1998) Molecular Cell 2: 361-372
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Holliday junction ATP-dependent DNA helicase RuvA | 203 | Mycobacterium leprae TN | Mutation(s): 0 Gene Names: ruvA, ML0482, B1177_C2_188 EC: 3.6.4.12 |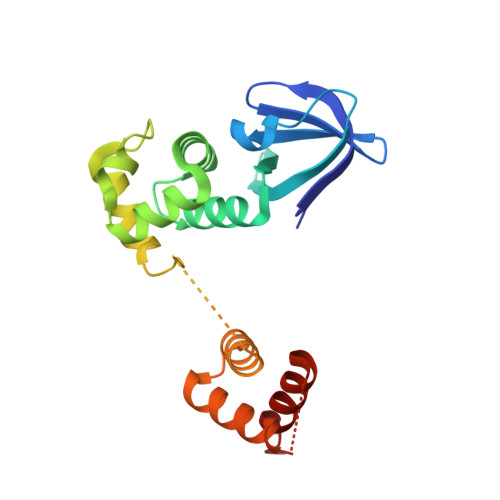  | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P40832 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*AP*GP*TP*TP*CP*GP*CP*GP*AP*GP*TP*TP*CP*GP*C)-3') | 15 | Mycobacterium leprae |  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 3 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*GP*CP*GP*AP*AP*CP*TP*CP*GP*CP*GP*AP*AP*C)-3') | 14 | Mycobacterium leprae |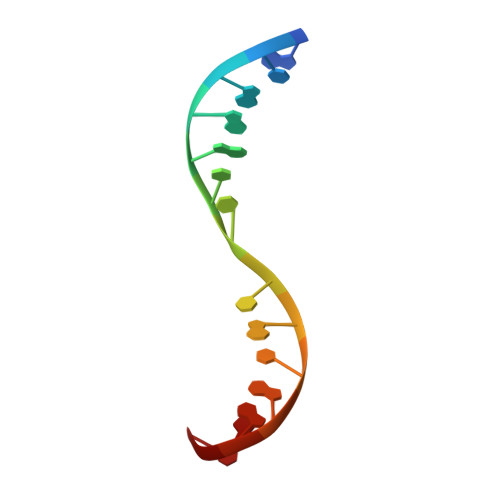  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 4 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(*AP*GP*TP*TP*CP*GP*CP*GP*CP*GP*CP*GP*AP*AP*CP*T)-3') | 16 | Mycobacterium leprae |  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 5 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(P*GP*TP*TP*CP*GP*CP*GP*CP*GP*CP*GP*AP*AP*CP*T)-3') | 15 | Mycobacterium leprae |  | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CA Download:Ideal Coordinates CCD File | M [auth K], N [auth L] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 141.35 | α = 90 |
| b = 141.35 | β = 90 |
| c = 106.47 | γ = 120 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| MOSFLM | data reduction |
| Aimless | data scaling |
| PHENIX | phasing |