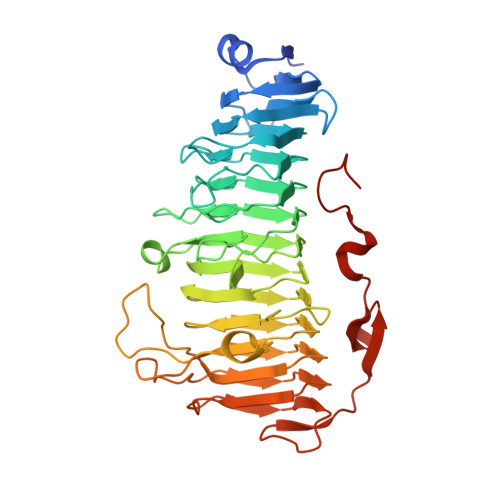
Exploring molecular determinants of polysaccharide lyase family 6-1 enzyme activity.
Violot, S., Galisson, F., Carrique, L., Jugnarain, V., Conchou, L., Robert, X., Thureau, A., Helbert, W., Aghajari, N., Ballut, L.(2021) Glycobiology 31: 1557-1570
- PubMed: 34245266 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwab073
- Primary Citation Related Structures:
7O77, 7O78, 7O79, 7O7A, 7O7T, 7O84 - PubMed Abstract:
The polysaccharide lyase family 6 (PL6) represents one of the 41 polysaccharide lyase families classified in the CAZy database with the vast majority of its members being alginate lyases grouped into three subfamilies, PL6_1-3. To decipher the mode of recognition and action of the enzymes belonging to subfamily PL6_1, we solved the crystal structures of Pedsa0632, Patl3640, Pedsa3628 and Pedsa3807, which all show different substrate specificities and mode of action (endo-/exolyase). Thorough exploration of the structures of Pedsa0632 and Patl3640 in complex with their substrates as well as docking experiments confirms that the conserved residues in subsites -1 to +3 of the catalytic site form a common platform that can accommodate various types of alginate in a very similar manner but with a series of original adaptations bringing them their specificities of action. From comparative studies with existing structures of PL6_1 alginate lyases, we observe that in the right-handed parallel β-helix fold shared by all these enzymes, the substrate-binding site harbors the same overall conserved structures and organization. Despite this apparent similarity, it appears that members of the PL6_1 subfamily specifically accommodate and catalyze the degradation of different alginates suggesting that this common platform is actually a highly adaptable and specific tool.
- Molecular Microbiology and Structural Biochemistry, UMR 5086, CNRS Université de Lyon, 7 passage du Vercors, Lyon 69367, France.
Organizational Affiliation: