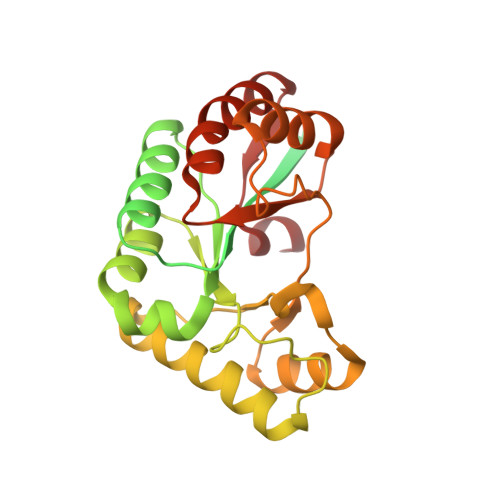
Crystal structure of acetylxylan esterase from Caldanaerobacter subterraneus subsp. tengcongensis.
Sasamoto, K., Himiyama, T., Moriyoshi, K., Ohmoto, T., Uegaki, K., Nishiya, Y., Nakamura, T.(2021) Acta Crystallogr F Struct Biol Commun 77: 399-406
- PubMed: 34726178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21009675
- Primary Citation Related Structures:
7FBW - PubMed Abstract:
The acetylxylan esterases (AXEs) classified into carbohydrate esterase family 4 (CE4) are metalloenzymes that catalyze the deacetylation of acetylated carbohydrates. AXE from Caldanaerobacter subterraneus subsp. tengcongensis (TTE0866), which belongs to CE4, is composed of three parts: a signal sequence (residues 1-22), an N-terminal region (NTR; residues 23-135) and a catalytic domain (residues 136-324). TTE0866 catalyzes the deacetylation of highly substituted cellulose acetate and is expected to be useful for industrial applications in the reuse of resources. In this study, the crystal structure of TTE0866 (residues 23-324) was successfully determined. The crystal diffracted to 1.9 Å resolution and belonged to space group I2 1 2 1 2 1 . The catalytic domain (residues 136-321) exhibited a (β/α) 7 -barrel topology. However, electron density was not observed for the NTR (residues 23-135). The crystal packing revealed the presence of an intermolecular space without observable electron density, indicating that the NTR occupies this space without a defined conformation or was truncated during the crystallization process. Although the active-site conformation of TTE0866 was found to be highly similar to those of other CE4 enzymes, the orientation of its Trp264 side chain near the active site was clearly distinct. The unique orientation of the Trp264 side chain formed a different-shaped cavity within TTE0866, which may contribute to its reactivity towards highly substituted cellulose acetate.
- Division of Life Science, Graduate School of Science and Engineering, Setsunan University, 17-8 Ikeda-Nakamachi, Neyagawa, Osaka 572-8508, Japan.
Organizational Affiliation: