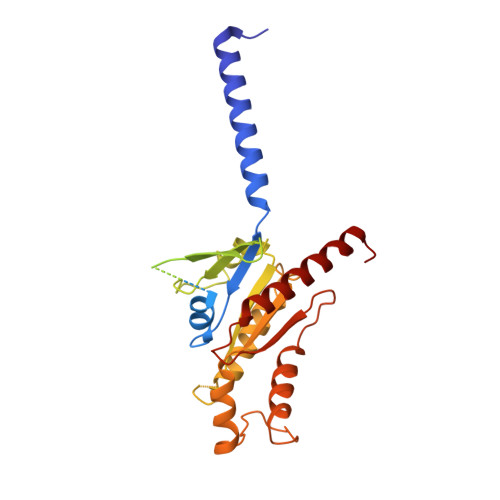
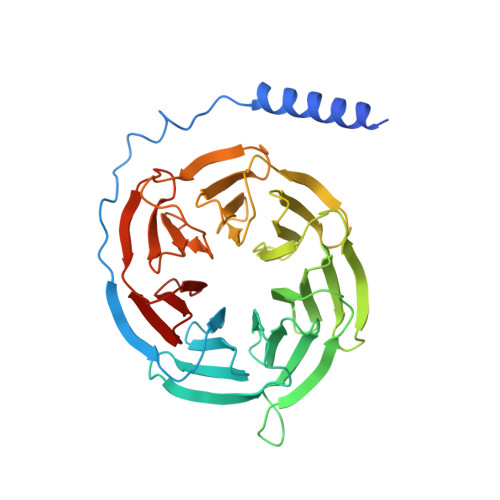
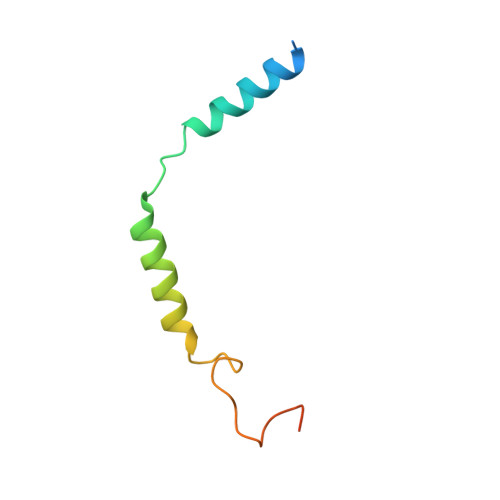
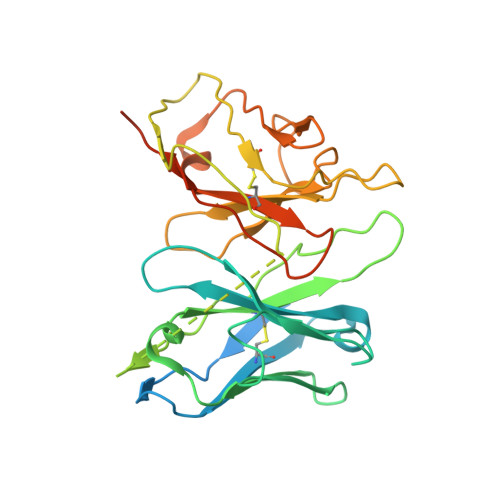
Structural insights into ligand recognition, activation, and signaling of the alpha 2A adrenergic receptor.
Xu, J., Cao, S., Hubner, H., Weikert, D., Chen, G., Lu, Q., Yuan, D., Gmeiner, P., Liu, Z., Du, Y.(2022) Sci Adv 8: eabj5347-eabj5347
- PubMed: 35245122 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abj5347
- Primary Citation Related Structures:
7EJ0, 7EJ8, 7EJA, 7EJK - PubMed Abstract:
The α 2A adrenergic receptor (α 2A AR) is a G protein (heterotrimeric guanine nucleotide-binding protein)-coupled receptor that mediates important physiological functions in response to the endogenous neurotransmitters norepinephrine and epinephrine, as well as numerous chemically distinct drugs. However, the molecular mechanisms of drug actions remain poorly understood. Here, we report the cryo-electron microscopy structures of the human α 2A AR-GoA complex bound to norepinephrine and three imidazoline derivatives (brimonidine, dexmedetomidine, and oxymetazoline). Together with mutagenesis and functional data, these structures provide important insights into the molecular basis of ligand recognition, activation, and signaling at the α 2A AR. Further structural analyses uncover different molecular determinants between α 2A AR and βARs for recognition of norepinephrine and key regions that determine the G protein coupling selectivity. Overall, our studies provide a framework for understanding the signal transduction of the adrenergic system at the atomic level, which will facilitate rational structure-based discovery of safer and more effective medications for α 2A AR.
- Kobilka Institute of Innovative Drug Discovery, School of Life and Health Sciences, Chinese University of Hong Kong, Shenzhen, 518172, China.
Organizational Affiliation: