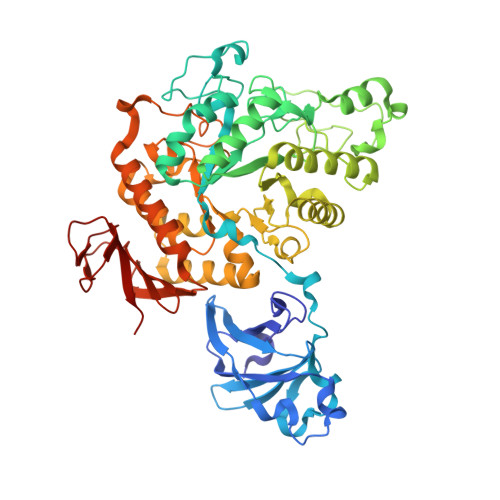
A GH13 alpha-glucosidase from Weissella cibaria uncommonly acts on short-chain maltooligosaccharides.
Wangpaiboon, K., Laohawuttichai, P., Kim, S.Y., Mori, T., Nakapong, S., Pichyangkura, R., Pongsawasdi, P., Hakoshima, T., Krusong, K.(2021) Acta Crystallogr D Struct Biol 77: 1064-1076
- PubMed: 34342279 Search on PubMed
- DOI: https://doi.org/10.1107/S205979832100677X
- Primary Citation Related Structures:
7D9B, 7D9C, 7DCG, 7DCH, 7EHH, 7EHI - PubMed Abstract:
α-Glucosidase (EC 3.2.1.20) is a carbohydrate-hydrolyzing enzyme which generally cleaves α-1,4-glycosidic bonds of oligosaccharides and starch from the nonreducing ends. In this study, the novel α-glucosidase from Weissella cibaria BBK-1 (WcAG) was biochemically and structurally characterized. WcAG belongs to glycoside hydrolase family 13 (GH13) and to the neopullanase subfamily. It exhibits distinct hydrolytic activity towards the α-1,4 linkages of short-chain oligosaccharides from the reducing end. The enzyme prefers to hydrolyse maltotriose and acarbose, while it cannot hydrolyse cyclic oligosaccharides and polysaccharides. In addition, WcAG can cleave pullulan hydrolysates and strongly exhibits transglycosylation activity in the presence of maltose. Size-exclusion chromatography and X-ray crystal structures revealed that WcAG forms a homodimer in which the N-terminal domain of one monomer is orientated in proximity to the catalytic domain of another, creating the substrate-binding groove. Crystal structures of WcAG in complexes with maltose, maltotriose and acarbose revealed a remarkable enzyme active site with accessible +2, +1 and -1 subsites, along with an Arg-Glu gate (Arg176-Glu296) in front of the active site. The -2 and -3 subsites were blocked by Met119 and Asn120 from the N-terminal domain of a different subunit, resulting in an extremely restricted substrate preference.
- Department of Biochemistry, Faculty of Science, Chulalongkorn University, Bangkok 10330, Thailand.
Organizational Affiliation: