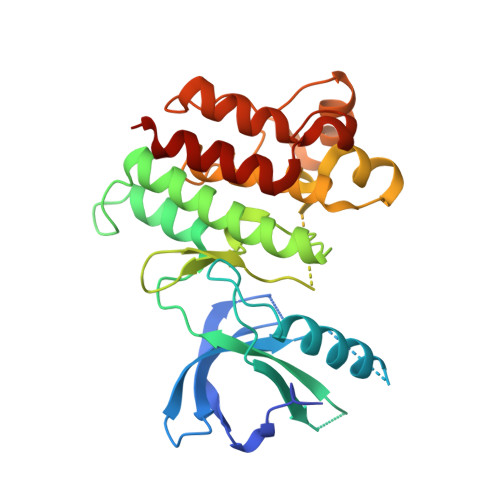
Crystal Structure of the Kinase Domain of MerTK in Complex with AZD7762 Provides Clues for Structure-Based Drug Development.
Park, T.H., Bae, S.H., Bong, S.M., Ryu, S.E., Jang, H., Lee, B.I.(2020) Int J Mol Sci 21
- PubMed: 33114206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms21217878
- Primary Citation Related Structures:
7CQE - PubMed Abstract:
Aberrant tyrosine-protein kinase Mer (MerTK) expression triggers prosurvival signaling and contributes to cell survival, invasive motility, and chemoresistance in many kinds of cancers. In addition, recent reports suggested that MerTK could be a primary target for abnormal platelet aggregation. Consequently, MerTK inhibitors may promote cancer cell death, sensitize cells to chemotherapy, and act as new antiplatelet agents. We screened an inhouse chemical library to discover novel small-molecule MerTK inhibitors, and identified AZD7762, which is known as a checkpoint-kinase (Chk) inhibitor. The inhibition of MerTK by AZD7762 was validated using an in vitro homogeneous time-resolved fluorescence (HTRF) assay and through monitoring the decrease in phosphorylated MerTK in two lung cancer cell lines. We also determined the crystal structure of the MerTK:AZD7762 complex and revealed the binding mode of AZD7762 to MerTK. Structural information from the MerTK:AZD7762 complex and its comparison with other MerTK:inhibitor structures gave us new insights for optimizing the development of inhibitors targeting MerTK.
- Research Institute, National Cancer Center, Goyang, 10408 Gyeonggi, Korea.
Organizational Affiliation: