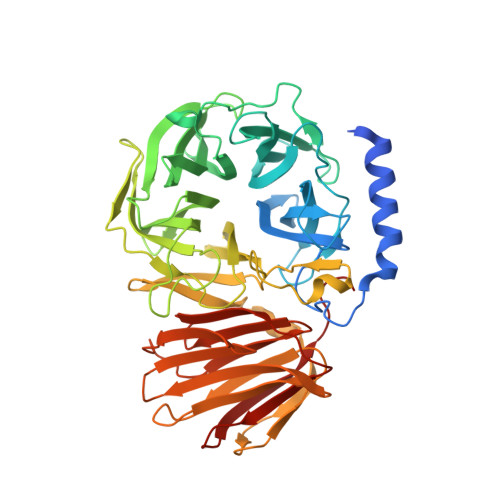
Structural insight into the substrate specificity of Bombyx mori beta-fructofuranosidase belonging to the glycoside hydrolase family 32.
Miyazaki, T., Oba, N., Park, E.Y.(2020) Insect Biochem Mol Biol 127: 103494-103494
- PubMed: 33132139 Search on PubMed
- DOI: https://doi.org/10.1016/j.ibmb.2020.103494
- Primary Citation Related Structures:
7BWB, 7BWC - PubMed Abstract:
Sucrose-hydrolyzing enzymes are largely divided into β-fructofuranosidase and sucrose α-glucosidase. The domestic silkworm Bombyx mori possesses both enzymes, BmSUC1 and BmSUH, belonging to the glycoside hydrolase family 32 (GH32) and GH13, respectively. BmSUC1 was presumed to be acquired by horizontal gene transfer from bacteria based on phylogenetic analysis and related to tolerance to sugar-mimic alkaloids contained in mulberry latex. Here we investigated the substrate specificity of recombinant BmSUC1 that can hydrolyze not only sucrose but also fructooligosaccharides and fructans, and revealed that the enzyme was competitively inhibited by 1,4-dideoxy-1,4-imino-D-arabinitol, one of the alkaloids. Moreover, the crystal structures of BmSUC1 in apo form and complex with sucrose were determined, and the active site pocket was shallow and suitable for shorter substrates but was related to more relaxed substrate specificity than the strict sucrose α-glucosidase BmSUH. Considering together with the distribution of BmSUC1-orthologous genes in many lepidopterans, our results suggest that BmSUC1 contributes to the digestion of fructooligosaccharides and fructans derived from feed plants.
- Green Chemistry Research Division, Research Institute of Green Science and Technology, Shizuoka University, 836 Ohya, Suruga-ku, Shizuoka, 422-8529, Japan; Department of Applied Life Sciences, Faculty of Agriculture, Shizuoka University, 836 Ohya, Suruga-ku, Shizuoka, 422-8529, Japan. Electronic address: miyazaki.takatsugu@shizuoka.ac.jp.
Organizational Affiliation: