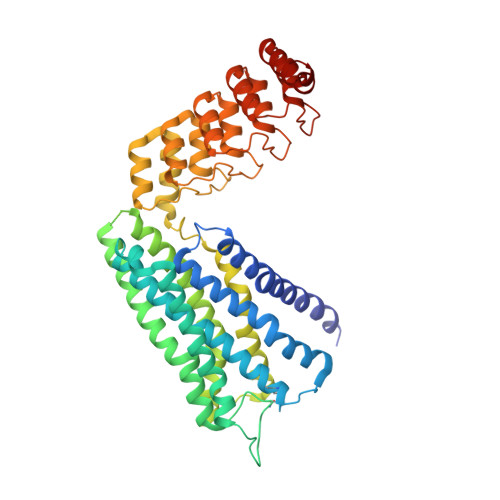
Crystal structure of the alpha 1B -adrenergic receptor reveals molecular determinants of selective ligand recognition.
Deluigi, M., Morstein, L., Schuster, M., Klenk, C., Merklinger, L., Cridge, R.R., de Zhang, L.A., Klipp, A., Vacca, S., Vaid, T.M., Mittl, P.R.E., Egloff, P., Eberle, S.A., Zerbe, O., Chalmers, D.K., Scott, D.J., Pluckthun, A.(2022) Nat Commun 13: 382-382
- PubMed: 35046410 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27911-3
- Primary Citation Related Structures:
7B6W - PubMed Abstract:
α-adrenergic receptors (αARs) are G protein-coupled receptors that regulate vital functions of the cardiovascular and nervous systems. The therapeutic potential of αARs, however, is largely unexploited and hampered by the scarcity of subtype-selective ligands. Moreover, several aminergic drugs either show off-target binding to αARs or fail to interact with the desired subtype. Here, we report the crystal structure of human α 1B AR bound to the inverse agonist (+)-cyclazosin, enabled by the fusion to a DARPin crystallization chaperone. The α 1B AR structure allows the identification of two unique secondary binding pockets. By structural comparison of α 1B AR with α 2 ARs, and by constructing α 1B AR-α 2C AR chimeras, we identify residues 3.29 and 6.55 as key determinants of ligand selectivity. Our findings provide a basis for discovery of α 1B AR-selective ligands and may guide the optimization of aminergic drugs to prevent off-target binding to αARs, or to elicit a selective interaction with the desired subtype.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057, Zurich, Switzerland.
Organizational Affiliation: