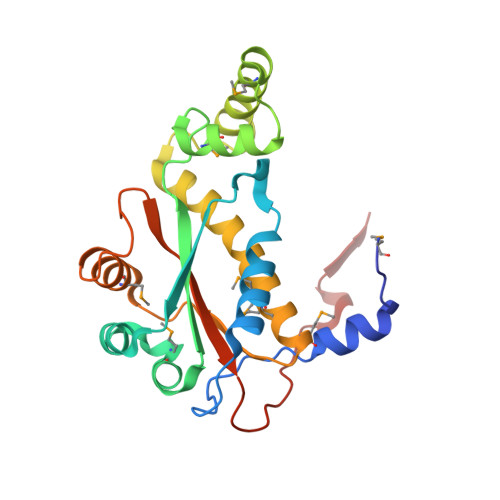
Functional and Structural Characterization of Diverse NfsB Chloramphenicol Reductase Enzymes from Human Pathogens.
Mullowney, M.W., Maltseva, N.I., Endres, M., Kim, Y., Joachimiak, A., Crofts, T.S.(2022) Microbiol Spectr 10: e0013922-e0013922
- PubMed: 35195438 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/spectrum.00139-22
- Primary Citation Related Structures:
6WT2, 7LDQ, 7RZL, 7RZP, 7S14, 7S1A - PubMed Abstract:
Phylogenetically diverse bacteria can carry out chloramphenicol reduction, but only a single enzyme has been described that efficiently catalyzes this reaction, the NfsB nitroreductase from Haemophilus influenzae strain KW20. Here, we tested the hypothesis that some NfsB homologs function as housekeeping enzymes with the potential to become chloramphenicol resistance enzymes. We found that expression of H. influenzae and Neisseria spp. nfsB genes, but not Pasteurella multocida nfsB , allows Escherichia coli to resist chloramphenicol by nitroreduction. Mass spectrometric analysis confirmed that purified H. influenzae and N. meningitides NfsB enzymes reduce chloramphenicol to amino-chloramphenicol, while kinetics analyses supported the hypothesis that chloramphenicol reduction is a secondary activity. We combined these findings with atomic resolution structures of multiple chloramphenicol-reducing NfsB enzymes to identify potential key substrate-binding pocket residues. Our work expands the chloramphenicol reductase family and provides mechanistic insights into how a housekeeping enzyme might confer antibiotic resistance. IMPORTANCE The question of how new enzyme activities evolve is of great biological interest and, in the context of antibiotic resistance, of great medical importance. Here, we have tested the hypothesis that new antibiotic resistance mechanisms may evolve from promiscuous housekeeping enzymes that have antibiotic modification side activities. Previous work identified a Haemophilus influenzae nitroreductase housekeeping enzyme that has the ability to give Escherichia coli resistance to the antibiotic chloramphenicol by nitroreduction. Herein, we extend this work to enzymes from other Haemophilus and Neisseria strains to discover that expression of chloramphenicol reductases is sufficient to confer chloramphenicol resistance to Es. coli, confirming that chloramphenicol reductase activity is widespread across this nitroreductase family. By solving the high-resolution crystal structures of active chloramphenicol reductases, we identified residues important for this activity. Our work supports the hypothesis that housekeeping proteins possessing multiple activities can evolve into antibiotic resistance enzymes.
- Department of Chemistry, Northwestern Universitygrid.16753.36, Evanston, Illinois, USA.
Organizational Affiliation: