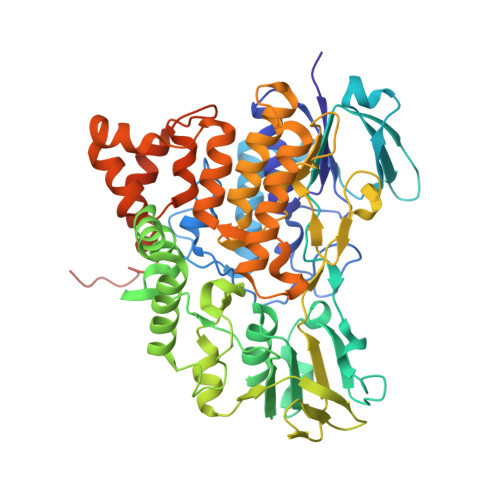
Ancestral-sequence reconstruction unveils the structural basis of function in mammalian FMOs.
Nicoll, C.R., Bailleul, G., Fiorentini, F., Mascotti, M.L., Fraaije, M.W., Mattevi, A.(2020) Nat Struct Mol Biol 27: 14-24
- PubMed: 31873300 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0347-2
- Primary Citation Related Structures:
6SE3, 6SEK, 6SEM, 6SF0 - PubMed Abstract:
Flavin-containing monooxygenases (FMOs) are ubiquitous in all domains of life and metabolize a myriad of xenobiotics, including toxins, pesticides and drugs. However, despite their pharmacological importance, structural information remains bereft. To further our understanding behind their biochemistry and diversity, we used ancestral-sequence reconstruction, kinetic and crystallographic techniques to scrutinize three ancient mammalian FMOs: AncFMO2, AncFMO3-6 and AncFMO5. Remarkably, all AncFMOs could be crystallized and were structurally resolved between 2.7- and 3.2-Å resolution. These crystal structures depict the unprecedented topology of mammalian FMOs. Each employs extensive membrane-binding features and intricate substrate-profiling tunnel networks through a conspicuous membrane-adhering insertion. Furthermore, a glutamate-histidine switch is speculated to induce the distinctive Baeyer-Villiger oxidation activity of FMO5. The AncFMOs exhibited catalysis akin to human FMOs and, with sequence identities between 82% and 92%, represent excellent models. Our study demonstrates the power of ancestral-sequence reconstruction as a strategy for the crystallization of proteins.
- Department of Biology and Biotechnology "Lazzaro Spallanzani", University of Pavia, Pavia, Italy.
Organizational Affiliation: