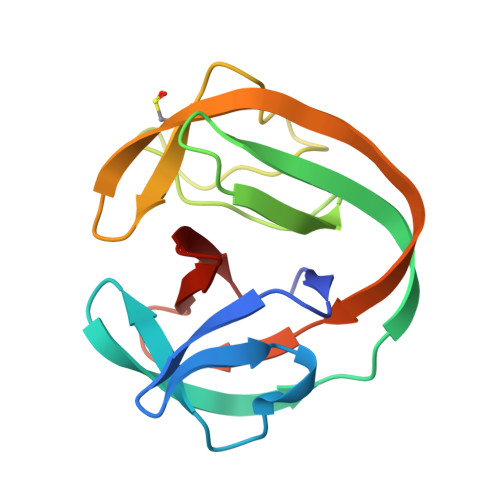
The Convergence of the Hedgehog/Intein Fold in Different Protein Splicing Mechanisms.
Beyer, H.M., Virtanen, S.I., Aranko, A.S., Mikula, K.M., Lountos, G.T., Wlodawer, A., Ollila, O.H.S., Iwai, H.(2020) Int J Mol Sci 21
- PubMed: 33171880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms21218367
- Primary Citation Related Structures:
6RIX, 6RIY, 6RIZ - PubMed Abstract:
Protein splicing catalyzed by inteins utilizes many different combinations of amino-acid types at active sites. Inteins have been classified into three classes based on their characteristic sequences. We investigated the structural basis of the protein splicing mechanism of class 3 inteins by determining crystal structures of variants of a class 3 intein from Mycobacterium chimaera and molecular dynamics simulations, which suggested that the class 3 intein utilizes a different splicing mechanism from that of class 1 and 2 inteins. The class 3 intein uses a bond cleavage strategy reminiscent of proteases but share the same Hedgehog/INTein (HINT) fold of other intein classes. Engineering of class 3 inteins from a class 1 intein indicated that a class 3 intein would unlikely evolve directly from a class 1 or 2 intein. The HINT fold appears as structural and functional solution for trans -peptidyl and trans -esterification reactions commonly exploited by diverse mechanisms using different combinations of amino-acid types for the active-site residues.
- Institute of Biotechnology, University of Helsinki, P.O. Box 65, FIN-00014 Helsinki, Finland.
Organizational Affiliation: