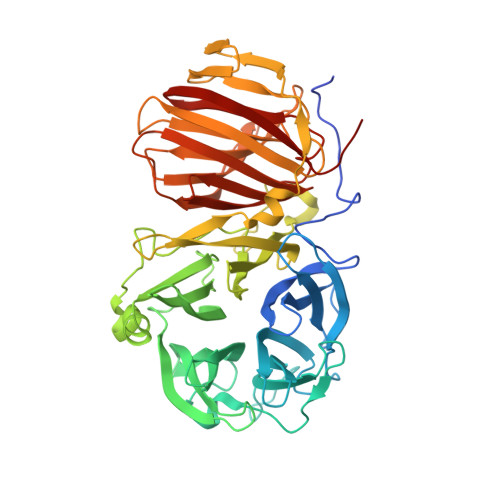
First crystal structure of an endo-levanase - the BT1760 from a human gut commensal Bacteroides thetaiotaomicron.
Ernits, K., Eek, P., Lukk, T., Visnapuu, T., Alamae, T.(2019) Sci Rep 9: 8443-8443
- PubMed: 31186460 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-44785-0
- Primary Citation Related Structures:
6R3R, 6R3U - PubMed Abstract:
The endo-levanase BT1760 of a human gut commensal Bacteroides thetaiotaomicron randomly cuts a β-2,6-linked fructan, levan, into fructo-oligosaccharides providing a prebiotic substrate for gut microbiota. Here we introduce the crystal structure of BT1760 at resolution of 1.65 Å. The fold of the enzyme is typical for GH32 family proteins: a catalytic N-terminal five-bladed β-propeller connected with a C-terminal β-sandwich domain. The levantetraose-bound structure of catalytically inactive mutant E221A at 1.90-Å resolution reveals differences in substrate binding between the endo-acting fructanases. A shallow substrate-binding pocket of the endo-inulinase INU2 of Aspergillus ficuum binds at least three fructose residues at its flat bottom. In the levantetraose-soaked crystal of the endo-levanase E221A mutant the ligand was bent into the pond-like substrate pocket with its fructose residues making contacts at -3, -2, -1 and + 1 subsites residing at several pocket depths. Binding of levantetraose to the β-sandwich domain was not detected. The N- and C-terminal modules of BT1760 did not bind levan if expressed separately, the catalytic domain lost its activity and both modules tended to precipitate. We gather that endo-levanase BT1760 requires both domains for correct folding, solubility and stability of the protein.
- Department of Genetics, Institute of Molecular and Cell Biology, University of Tartu, Riia 23, 51010, Tartu, Estonia.
Organizational Affiliation: