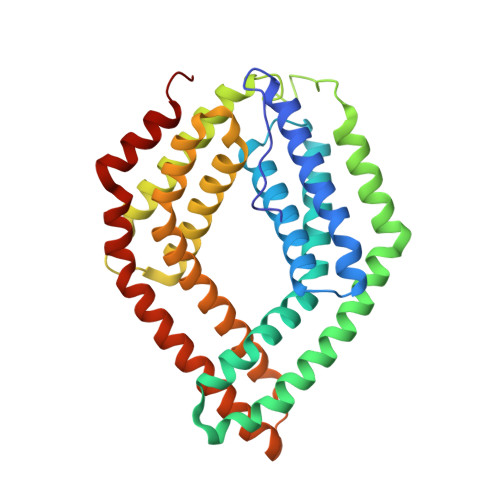
Molecular characterization of CHAD domains as inorganic polyphosphate-binding modules.
Lorenzo-Orts, L., Hohmann, U., Zhu, J., Hothorn, M.(2019) Life Sci Alliance 2
- PubMed: 31133615 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.201900385
- Primary Citation Related Structures:
6QV5, 6QV7, 6QVA - PubMed Abstract:
Inorganic polyphosphates (polyPs) are linear polymers of orthophosphate units linked by phosphoanhydride bonds. Here, we report that bacterial, archaeal, and eukaryotic conserved histidine α-helical (CHAD) domains are specific polyP-binding modules. Crystal structures reveal that CHAD domains are formed by two four-helix bundles, giving rise to a central pore surrounded by conserved basic surface patches. Different CHAD domains bind polyPs with dissociation constants ranging from the nano- to mid-micromolar range, but not nucleic acids. A CHAD-polyP complex structure reveals the phosphate polymer binding across the central pore and along the two basic patches. Mutational analysis of CHAD-polyP interface residues validates the complex structure. The presence of a CHAD domain in the polyPase ygiF enhances its enzymatic activity. The only known CHAD protein from the plant Ricinus communis localizes to the nucleus/nucleolus when expressed in Arabidopsis and tobacco, suggesting that plants may harbor polyPs in these compartments. We propose that CHAD domains may be used to engineer the properties of polyP-metabolizing enzymes and to specifically localize polyP stores in eukaryotic cells and tissues.
- Structural Plant Biology Laboratory, Department of Botany and Plant Biology, Faculty of Sciences, University of Geneva, Geneva, Switzerland.
Organizational Affiliation: