Biomimetic Macrocyclic Inhibitors of Human Cathepsin D: Structure-Activity Relationship and Binding Mode Analysis.
Houstecka, R., Hadzima, M., Fanfrlik, J., Brynda, J., Pallova, L., Hanova, I., Mertlikova-Kaiserova, H., Lepsik, M., Horn, M., Smrcina, M., Majer, P., Mares, M.(2020) J Med Chem 63: 1576-1596
- PubMed: 32003991 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01351
- Primary Citation Related Structures:
6QBG, 6QBH, 6QCB - PubMed Abstract:
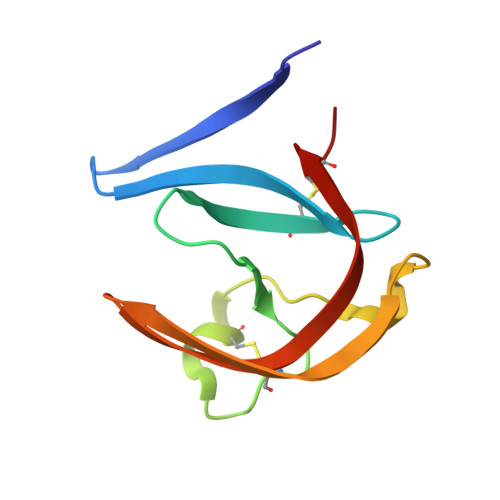
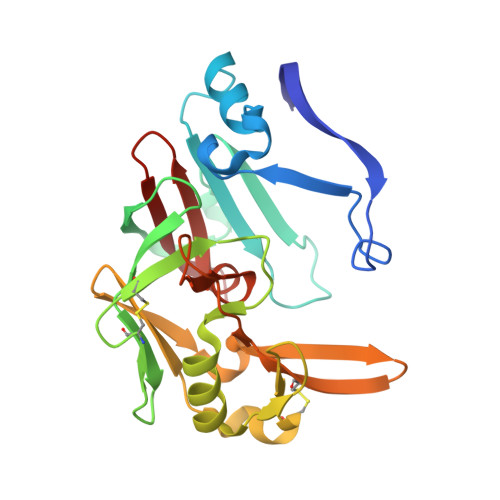
Human cathepsin D (CatD), a pepsin-family aspartic protease, plays an important role in tumor progression and metastasis. Here, we report the development of biomimetic inhibitors of CatD as novel tools for regulation of this therapeutic target. We designed a macrocyclic scaffold to mimic the spatial conformation of the minimal pseudo-dipeptide binding motif of pepstatin A, a microbial oligopeptide inhibitor, in the CatD active site. A library of more than 30 macrocyclic peptidomimetic inhibitors was employed for scaffold optimization, mapping of subsite interactions, and profiling of inhibitor selectivity. Furthermore, we solved high-resolution crystal structures of three macrocyclic inhibitors with low nanomolar or subnanomolar potency in complex with CatD and determined their binding mode using quantum chemical calculations. The study provides a new structural template and functional profile that can be exploited for design of potential chemotherapeutics that specifically inhibit CatD and related aspartic proteases.
- Institute of Organic Chemistry and Biochemistry , Czech Academy of Sciences , Flemingovo nám. 2 , 16610 Praha 6 , Czech Republic.
Organizational Affiliation: